
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 13,767,086 – 13,767,259 |
| Length | 173 |
| Max. P | 0.762194 |

| Location | 13,767,086 – 13,767,179 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 48.00 |
| Shannon entropy | 1.09343 |
| G+C content | 0.44371 |
| Mean single sequence MFE | -27.51 |
| Consensus MFE | -5.31 |
| Energy contribution | -6.58 |
| Covariance contribution | 1.27 |
| Combinations/Pair | 2.07 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.19 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.711529 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 13767086 93 + 21146708 ---------UUGGUUUUCAUUGAAGGACAUCUUUAAGUCCAAUGA---------AAGUUUAGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCG ---------........(((((((((((........))))...((---------(.(((.((....(((((((......))))))).....))))).)))..))))))).. ( -25.30, z-score = -2.61, R) >droEre2.scaffold_4845 7994789 102 + 22589142 ---------UUGUUGCCCAUUGCGGGGCAGCUUUGGGGCCACAGUCAGUUGAGUAAGUUGGGUAAUUGGCUCCUUAUUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCG ---------..(((((((......))))))).....(((....))).((((((((.((((((....(((((((......)))))))....))))))....))))))))... ( -35.80, z-score = -1.61, R) >droYak2.chr2R 5723710 93 - 21139217 ---------UUGUUGUCCAUUGUGGGGCAACUUUAAGUCCACAGU---------AAGUUGAGUAGUUGGCUCUUUGCUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCG ---------..(((((((......))))))).....(((.((((.---------..(((((((.((.((((((......)))))))).)))).)))...))))..)))... ( -31.40, z-score = -2.77, R) >droSec1.super_1 11262660 93 + 14215200 ---------UUGGGUUUCGUUGAAGGACAACUUUAAGUCCACUGG---------AAGUUUGGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCG ---------........(((((((((((........))))...((---------(((((.((....(((((((......)))))))....)).))).)))).))))))).. ( -26.30, z-score = -1.72, R) >droSim1.chr2R 12508966 93 + 19596830 ---------UUGGUUUUCGUUAAAGGACAUCUUUAAGUCCACUGG---------AAGUUUAGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCG ---------...((..(((.....((((........))))...((---------((..((((....(((((((......))))))).....))))..))))...))).)). ( -23.40, z-score = -1.71, R) >droWil1.scaffold_180745 2230479 81 + 2843958 ---------------------------UUCCUGAAAGAACGUUGAUGCU---GCGGAUGAUGCUUUUCUUUUAUGGAAUGGAAUGACAUGGUCAAUGUUUUGCCAAAUGCC ---------------------------((((((((((((.((....)).---(((.....)))..)))))))).))))(((...(((((.....)))))...)))...... ( -18.20, z-score = -0.96, R) >droVir3.scaffold_12875 5200638 107 - 20611582 ----AAAAUUUCAUUGGCAGUACGCCUUUUACGGGGCAGCAGAAAUUUCCAAAAGGAUUCUGUUCUUUUAUCUUCAAUAUUUUUCAAAAACUCGGCAUUCUGUUCGAUUCG ----........((((((((...((((......))))(((((((...(((....))))))))))...................................))).)))))... ( -18.10, z-score = -0.59, R) >droMoj3.scaffold_6496 8382322 111 + 26866924 AACUAUUUGCUCGCCCUUCGUCUGCUGCUUGCUCUGGCUGACGUACUUGAGAAGGGAUGCUGCACGUUUCUCCUCGAUGCUGCCAUCGAUGUUCGCUUUCUGGUCGACAAU .......(((.(((((((((((.((.....))...)))...((....)).)))))).))..))).........((((((....))))))((((.(((....))).)))).. ( -26.10, z-score = 0.68, R) >droGri2.scaffold_15245 6109255 111 - 18325388 UUCAGUUGUUUCAGGGGAGGAAGGCUGCCUGUUGUGGGUCAAGAGGUUGAGGAACGAGGCAGCCUUCUUAUCCUUGAUCUGUUGCACAAAGUCAGCAUGCUGCUCGAUGCG ..(((.....(((((((((((((((((((((((.(...((((....))))).))).))))))))))))).))))))).)))..(((...((.(((....)))))...))). ( -43.00, z-score = -1.99, R) >consensus _________UUGGUUUUCAUUGAGGGGCAUCUUUAAGUCCACUGA_________AAGUUCAGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCG ........................((((........))))..........................((((((........))))))........(((((......))))). ( -5.31 = -6.58 + 1.27)


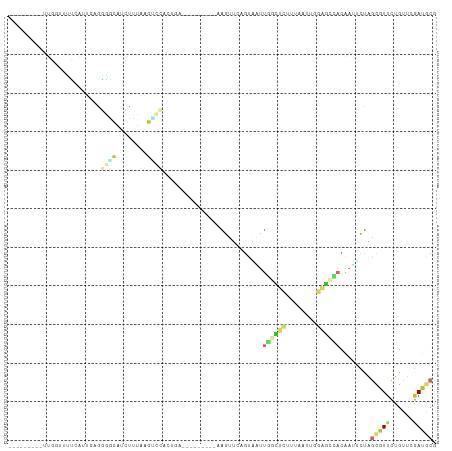
| Location | 13,767,112 – 13,767,219 |
|---|---|
| Length | 107 |
| Sequences | 10 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 57.58 |
| Shannon entropy | 0.89014 |
| G+C content | 0.49909 |
| Mean single sequence MFE | -33.10 |
| Consensus MFE | -10.88 |
| Energy contribution | -9.97 |
| Covariance contribution | -0.91 |
| Combinations/Pair | 2.09 |
| Mean z-score | -1.04 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.762194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 13767112 107 + 21146708 -------AGUCCAAUGA--AAGUUUAGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCCGUCAGUCGCUCCUGACAGCCGCUAGCA -------..........--............(((((((......))))))).....((((((..(((((.(..(((.(((((......)).))).)))..).))))).)))))).. ( -34.60, z-score = -2.69, R) >droAna3.scaffold_13337 4134837 100 + 23293914 -------AGUCUGCGGA---------GUCAUUGGGUUGCCCUUUGAAGCCAAGAGUCUAGCGUGUUGCUGGACUGUAUUGCGAAUGUUCGUCUGUCGCUCCCGGCAAUCGUUGGCA -------.(((.((((.---------.(((..(((...)))..))).(((.(.((((((((.....)))))))).)...((((..(.....)..))))....)))..)))).))). ( -34.20, z-score = -1.12, R) >droEre2.scaffold_4845 7994815 116 + 22589142 GGGCCACAGUCAGUUGAGUAAGUUGGGUAAUUGGCUCCUUAUUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGCCCGUCAGUCGCUCCUGACAGCCGUUAGCA (((((.(((.(.((((((((.((((((....(((((((......)))))))....))))))....))))))))..).))).)...))))(((((......))))).((.....)). ( -40.20, z-score = -1.56, R) >droYak2.chr2R 5723736 107 - 21139217 -------AGUCCACAGU--AAGUUGAGUAGUUGGCUCUUUGCUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGCCCGUCAGUCGCUCCUGACAGCCGUUAGCA -------.(((.((((.--..(((((((.((.((((((......)))))))).)))).)))...))))..)))(((.(((((......)).))).)))..(((((....))))).. ( -33.80, z-score = -1.07, R) >droSec1.super_1 11262686 107 + 14215200 -------AGUCCACUGG--AAGUUUGGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCCGUCAGUCGCUCCUGACAGCCGCUAGCA -------.....((..(--....)..))...(((((((......))))))).....((((((..(((((.(..(((.(((((......)).))).)))..).))))).)))))).. ( -35.50, z-score = -1.81, R) >droSim1.chr2R 12508992 107 + 19596830 -------AGUCCACUGG--AAGUUUAGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCAGUCAGUCGCUCUUGACAGCCGCUAGCA -------.....(((((--....)))))...(((((((......))))))).....((((((..(((((.((.(((.((((........).))).)))))..))))).)))))).. ( -32.80, z-score = -1.25, R) >droWil1.scaffold_180745 2230496 104 + 2843958 -------AUGCUGCGGA-----UGAUGCUUUUCUUUUAUGGAAUGGAAUGACAUGGUCAAUGUUUUGCCAAAUGCCUUCACGUAUAUUACGCAAACGCGUUUUGCAAUCACUGGCA -------.((((...((-----(((.((..((((.....))))(((...(((((.....)))))...)))...))..))..(((....((((....))))..))).)))...)))) ( -22.10, z-score = 0.85, R) >droVir3.scaffold_12875 5200676 109 - 20611582 -------AAAUUUCCAAAAGGAUUCUGUUCUUUUAUCUUCAAUAUUUUUCAAAAACUCGGCAUUCUGUUCGAUUCGUUGGAGUAUGUCCUGUUGUUGCCUCUUGUAGAUGUCGGCC -------......((((((((((...))))))))((((.(((.......(((.(((..(((((....(((((....)))))..)))))..))).)))....))).))))...)).. ( -15.40, z-score = 0.85, R) >droMoj3.scaffold_6496 8382362 111 + 26866924 -----ACGUACUUGAGAAGGGAUGCUGCACGUUUCUCCUCGAUGCUGCCAUCGAUGUUCGCUUUCUGGUCGACAAUCUGGCGCAGCACGUGCUGGCGCUUCUUGUUGUCGCUGGCG -----..(((.(((((.((..(((.....)))..)).)))))))).((((.(((.....(((....)))((((((...((((((((....))).)))))..))))))))).)))). ( -38.80, z-score = -0.58, R) >droGri2.scaffold_15245 6109295 111 - 18325388 -----AAGAGGUUGAGGAACGAGGCAGCCUUCUUAUCCUUGAUCUGUUGCACAAAGUCAGCAUGCUGCUCGAUGCGACGACGUAUGUCCAGUUGCCGCUUCUUGCAGCCGCUGGCA -----((.((((..((((..((((....))))...))))..)))).)).......((((((..(((((..((.(((.((((.........)))).))).))..))))).)))))). ( -43.60, z-score = -2.02, R) >consensus _______AGUCUACUGA__AAGUUCAGUAAUUGGCUCUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCCGUCAGUCGCUCCUGACAGCCGCUAGCA ...............................((((((........)))))).....((((((..((((...........((((.((.....)).)))).....)))).)))))).. (-10.88 = -9.97 + -0.91)

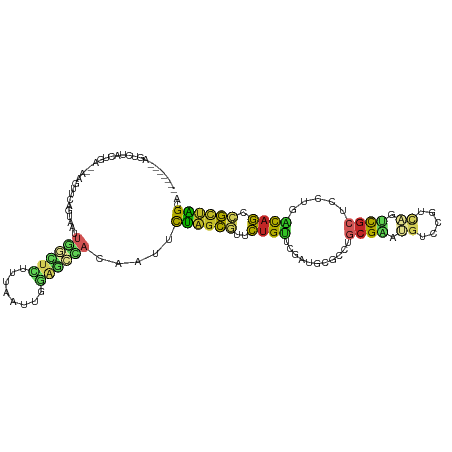

| Location | 13,767,139 – 13,767,259 |
|---|---|
| Length | 120 |
| Sequences | 10 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 64.87 |
| Shannon entropy | 0.76906 |
| G+C content | 0.50743 |
| Mean single sequence MFE | -37.03 |
| Consensus MFE | -7.62 |
| Energy contribution | -6.39 |
| Covariance contribution | -1.23 |
| Combinations/Pair | 2.00 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.21 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.719379 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 13767139 120 + 21146708 CUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCCGUCAGUCGCUCCUGACAGCCGCUAGCAUAGUAGCUCCAAGUGGCACGGAAAACCGAAUCCCUAUUUG .......((((((.((.((.((((((..(((((.(..(((.(((((......)).))).)))..).))))).)))))).)).)).))))))((.((..(((....)))...))))..... ( -41.20, z-score = -3.52, R) >droAna3.scaffold_13337 4134857 120 + 23293914 UGCCCUUUGAAGCCAAGAGUCUAGCGUGUUGCUGGACUGUAUUGCGAAUGUUCGUCUGUCGCUCCCGGCAAUCGUUGGCAUAGUAGCUCCAGGAGGCCCGGAACAGGCCCUCCAUGUUGG ((((...(((.(((.(.((((((((.....)))))))).)...((((..(.....)..))))....)))..)))..))))...((((....(((((.((......)).)))))..)))). ( -39.60, z-score = -0.31, R) >droEre2.scaffold_4845 7994851 120 + 22589142 CCUUAUUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGCCCGUCAGUCGCUCCUGACAGCCGUUAGCAUAGUAGCUCCAUGUGGCCCGGAACAGCGAAUCCCUACUUG .......((((((.((.((.((((((..(((((.(..(((.(((((......)).))).)))..).))))).)))))).)).)).)))))).((((...(((........)))))))... ( -38.00, z-score = -2.11, R) >droYak2.chr2R 5723763 120 - 21139217 CUUUGCUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGCCCGUCAGUCGCUCCUGACAGCCGUUAGCAUAGUAGCUCCAAGUGGCCCGGAACACCGAGUCCCUGUUUG ....(((((((((.((.((.((((((..(((((.(..(((.(((((......)).))).)))..).))))).)))))).)).)).)))))))))(((.(((....))).)))........ ( -45.60, z-score = -3.82, R) >droSec1.super_1 11262713 120 + 14215200 CUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCCGUCAGUCGCUCCUGACAGCCGCUAGCAUAGUAGCUCCAAGUGGCCCGGAAAACCGAAUCCCUAUUUG .......((((((.((.((.((((((..(((((.(..(((.(((((......)).))).)))..).))))).)))))).)).)).))))))((.((..(((....)))...))))..... ( -41.20, z-score = -3.77, R) >droSim1.chr2R 12509019 120 + 19596830 CUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCAGUCAGUCGCUCUUGACAGCCGCUAGCAUAGUAGCUCCAAGUGGCCCGGAAAACCGAAUCCCUAUUGG .......((((((.((.((.((((((..(((((.((.(((.((((........).))).)))))..))))).)))))).)).)).))))))((.((..(((....)))...))))..... ( -38.40, z-score = -2.19, R) >droWil1.scaffold_180745 2230520 113 + 2843958 UAUGGAAUGGAAUGACAUGGUCAAUGUUUUGCCAAAUGCCUUCACGUAUAUUACGCAAACGCGUUUUGCAAUCACUGGCAUAGUAACUCCAUGAGUCACGAAAGAAAGCAUCC------- ...((((((((...((...((((.((..((((.((((((.((..((((...))))..)).)))))).)))).)).))))...))...)))))......(....)......)))------- ( -27.20, z-score = -0.87, R) >droVir3.scaffold_12875 5200705 120 - 20611582 CUUCAAUAUUUUUCAAAAACUCGGCAUUCUGUUCGAUUCGUUGGAGUAUGUCCUGUUGUUGCCUCUUGUAGAUGUCGGCCAAGUAGCUCCAAUAGGCACAAAAUACUUCAUUAUUAUUUC ....................(((((.....).)))).....(((((((((((((((((..((..((((..(....)...))))..))..))))))).)))...))))))).......... ( -25.60, z-score = -1.70, R) >droMoj3.scaffold_6496 8382393 120 + 26866924 CCUCGAUGCUGCCAUCGAUGUUCGCUUUCUGGUCGACAAUCUGGCGCAGCACGUGCUGGCGCUUCUUGUUGUCGCUGGCGAAAUAGUUCCAAUACUCACAGAAAUCCUUGUUCAAAUUAU ..((((((....))))))..((((((....((.(((((((..((((((((....))).)))))....)))))))))))))))...................................... ( -33.50, z-score = -0.92, R) >droGri2.scaffold_15245 6109326 120 - 18325388 CCUUGAUCUGUUGCACAAAGUCAGCAUGCUGCUCGAUGCGACGACGUAUGUCCAGUUGCCGCUUCUUGCAGCCGCUGGCAAAGUAACUCCAAUAGGCCCGGAACAGUGAGUCCGUGUCAU .........(((((.....((((((..(((((..((.(((.((((.........)))).))).))..))))).))))))...))))).......(((.((((........)))).))).. ( -40.00, z-score = -1.35, R) >consensus CUUUAAUUGGAGCCACAAUUCUAGCGUUCUGUUCGAUGCGCCUGCGAAUGUCCGUCAGUCGCUCCUGACAGCCGCUAGCAUAGUAGCUCCAAGAGGCCCGGAAAACCGAAUCCCUAUUUG ....................((((((..((((...........((((.((.....)).)))).....)))).)))))).......................................... ( -7.62 = -6.39 + -1.23)

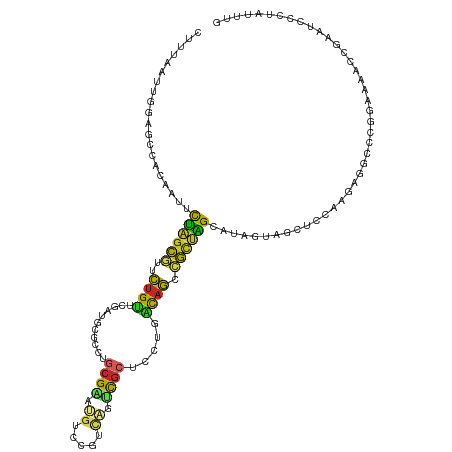
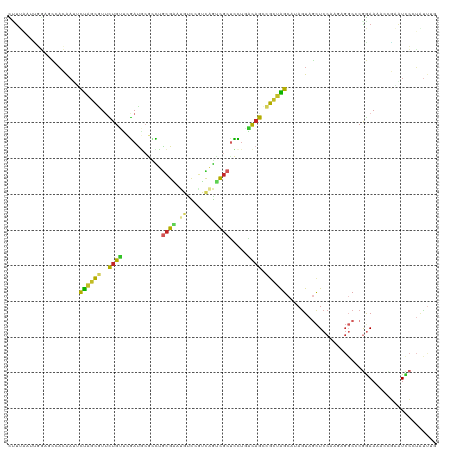
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:30:46 2011