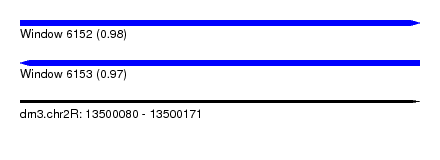

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 13,500,080 – 13,500,171 |
| Length | 91 |
| Max. P | 0.976535 |

| Location | 13,500,080 – 13,500,171 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 79.01 |
| Shannon entropy | 0.39544 |
| G+C content | 0.49145 |
| Mean single sequence MFE | -22.04 |
| Consensus MFE | -10.57 |
| Energy contribution | -11.93 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.89 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.95 |
| SVM RNA-class probability | 0.976535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
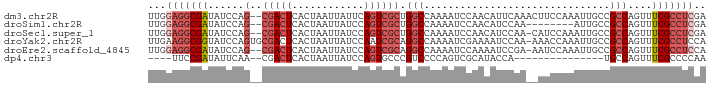
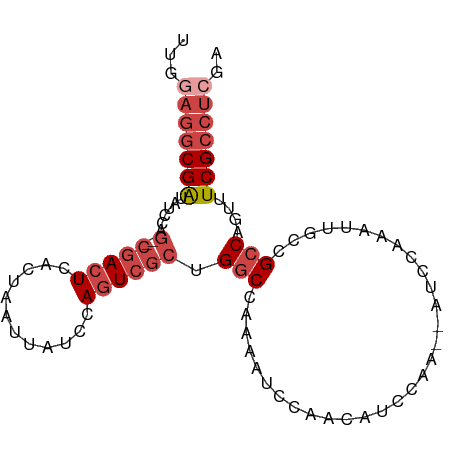
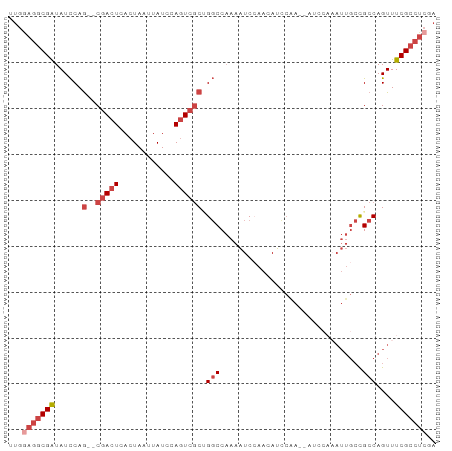
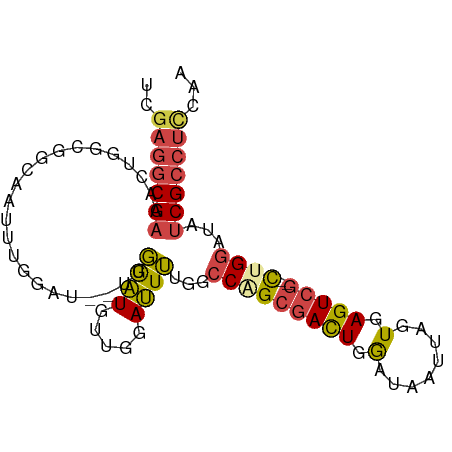
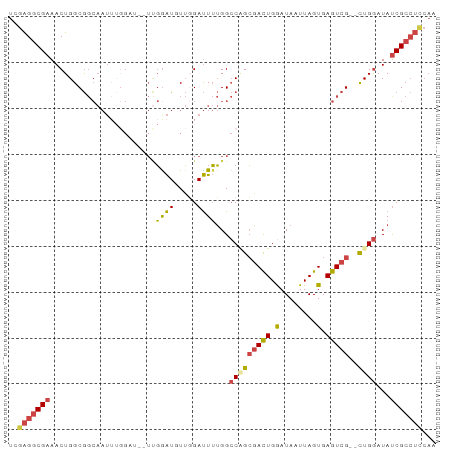
>dm3.chr2R 13500080 91 + 21146708 UUGGAGGCGAUAUCCAG--CGACUCACUAAUUAUUCAGUCGCUGGCCAAAAUCCAACAUUCAAACUUCCAAAUUGCCGCCAGUUUCGCCUCGA ...(((((((...((((--(((((............))))))))).........................(((((....)))))))))))).. ( -27.30, z-score = -3.96, R) >droSim1.chr2R 12237579 83 + 19596830 UUGGAGGCGAUAUCCAG--CGACUCACUAAUUAUCCAGUCGCUGGCCAAAAUCCAACAUCCAA--------AUUGCCGCCAGUUUCGCCUCGA ...(((((((...((((--(((((............))))))))).................(--------((((....)))))))))))).. ( -27.30, z-score = -3.94, R) >droSec1.super_1 10999180 90 + 14215200 UUGGAGGCGAUAUCCAG--CGACUCACUAAUUAUCCAGUCGCUGGCCAAAAUCCAACAUCCAA-CAUCCAAAUUGCCGCCAGUUUCGCCUCGA ...(((((((...((((--(((((............)))))))))..................-......(((((....)))))))))))).. ( -27.30, z-score = -3.96, R) >droYak2.chr2R 5453563 92 - 21139217 UUGAAGGCGGUAUCCAGUGCGACUCACUAAUUAUCCAAUCGCAGGCCAAAAUCGAAAAUCCAA-AAACCAAAUUGCCGCCAGUUUCGCCUCCA .....(((((((.....(((((................)))))((..................-...))....)))))))............. ( -16.39, z-score = -0.35, R) >droEre2.scaffold_4845 7728655 90 + 22589142 UUGGAGGCGAUAUCCAG--CGACUCACUAAUUAUCCAGUCGCAGGCCAAAAUCCAAAAUCCGA-AAUCCAAAUUGCCGCCAGUUUCGCCUCCA .(((((((((......(--(((((............)))))).(((...(((...........-.......)))...)))....))))))))) ( -26.67, z-score = -4.07, R) >dp4.chr3 9066970 72 - 19779522 ----UUCCGAUAUUCAA--CGACUCACUAAUUAUCCAGUGCCCGUCCCCAGUCGCAUACCA---------------UGCCAGUUUCGCCCCAA ----...(((.(((...--.((((((((........)))).........))))((((...)---------------))).))).)))...... ( -7.30, z-score = -1.04, R) >consensus UUGGAGGCGAUAUCCAG__CGACUCACUAAUUAUCCAGUCGCUGGCCAAAAUCCAACAUCCAA__AUCCAAAUUGCCGCCAGUUUCGCCUCGA ...(((((((.........(((((............)))))..(((...............................)))....))))))).. (-10.57 = -11.93 + 1.36)
| Location | 13,500,080 – 13,500,171 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 79.01 |
| Shannon entropy | 0.39544 |
| G+C content | 0.49145 |
| Mean single sequence MFE | -30.55 |
| Consensus MFE | -15.29 |
| Energy contribution | -17.82 |
| Covariance contribution | 2.53 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.75 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.971886 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
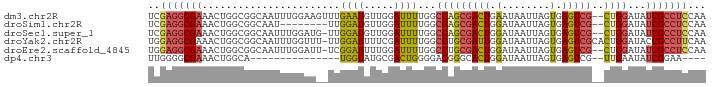
>dm3.chr2R 13500080 91 - 21146708 UCGAGGCGAAACUGGCGGCAAUUUGGAAGUUUGAAUGUUGGAUUUUGGCCAGCGACUGAAUAAUUAGUGAGUCG--CUGGAUAUCGCCUCCAA ..(((((((..(..(((.(((.........)))..)))..).......(((((((((.(........).)))))--))))...)))))))... ( -33.70, z-score = -3.32, R) >droSim1.chr2R 12237579 83 - 19596830 UCGAGGCGAAACUGGCGGCAAU--------UUGGAUGUUGGAUUUUGGCCAGCGACUGGAUAAUUAGUGAGUCG--CUGGAUAUCGCCUCCAA ..(((((((..(..(((.(...--------...).)))..).......(((((((((.(........).)))))--))))...)))))))... ( -33.40, z-score = -3.40, R) >droSec1.super_1 10999180 90 - 14215200 UCGAGGCGAAACUGGCGGCAAUUUGGAUG-UUGGAUGUUGGAUUUUGGCCAGCGACUGGAUAAUUAGUGAGUCG--CUGGAUAUCGCCUCCAA ..(((((((..(..(((.((((......)-)))..)))..).......(((((((((.(........).)))))--))))...)))))))... ( -36.70, z-score = -3.83, R) >droYak2.chr2R 5453563 92 + 21139217 UGGAGGCGAAACUGGCGGCAAUUUGGUUU-UUGGAUUUUCGAUUUUGGCCUGCGAUUGGAUAAUUAGUGAGUCGCACUGGAUACCGCCUUCAA ((((((((...(..(.(((...((((...-........)))).....)))(((((((.(........).))))))))..)....)))))))). ( -26.40, z-score = -0.96, R) >droEre2.scaffold_4845 7728655 90 - 22589142 UGGAGGCGAAACUGGCGGCAAUUUGGAUU-UCGGAUUUUGGAUUUUGGCCUGCGACUGGAUAAUUAGUGAGUCG--CUGGAUAUCGCCUCCAA (((((((((....(((.(.((((..((..-......))..)))).).))).((((((.(........).)))))--)......))))))))). ( -36.70, z-score = -4.10, R) >dp4.chr3 9066970 72 + 19779522 UUGGGGCGAAACUGGCA---------------UGGUAUGCGACUGGGGACGGGCACUGGAUAAUUAGUGAGUCG--UUGAAUAUCGGAA---- .....((.......)).---------------(((((((((((((....)...((((((....)))))))))))--)...))))))...---- ( -16.40, z-score = -0.87, R) >consensus UCGAGGCGAAACUGGCGGCAAUUUGGAU__UUGGAUGUUGGAUUUUGGCCAGCGACUGGAUAAUUAGUGAGUCG__CUGGAUAUCGCCUCCAA ..(((((((..(((((.(.((((..(...........)..)))).).)))))(((((.(........).))))).........)))))))... (-15.29 = -17.82 + 2.53)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:30:11 2011