
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 13,478,948 – 13,479,044 |
| Length | 96 |
| Max. P | 0.637183 |

| Location | 13,478,948 – 13,479,044 |
|---|---|
| Length | 96 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 67.55 |
| Shannon entropy | 0.66123 |
| G+C content | 0.53421 |
| Mean single sequence MFE | -22.13 |
| Consensus MFE | -7.26 |
| Energy contribution | -7.01 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637183 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
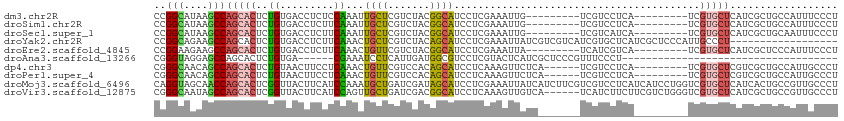
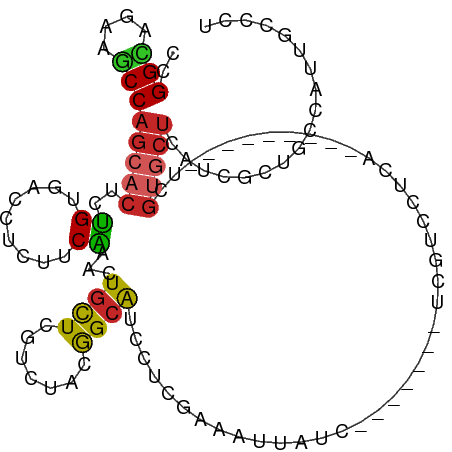
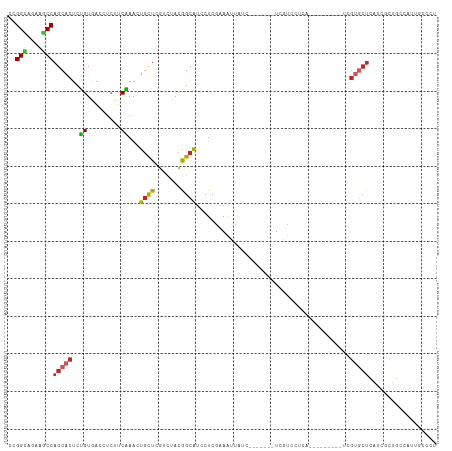
>dm3.chr2R 13478948 96 - 21146708 CCGGCAUAAGCCAGCACUCUGUGACCUCUCCAAAUUGCUCGUCUACGGCAUCCUCGAAAUUG---------UCGUCCUCA---------UCGUGCUCAUCGCUGCCAUUUCCCU ..((((......(((((..((.(((..(((.....(((.((....))))).....))....)---------..)))..))---------..)))))......))))........ ( -19.80, z-score = -1.38, R) >droSim1.chr2R 12214881 96 - 19596830 CCGGCAUAAGCCAGCACUCUGUGACCUCUUCAAAUUGCUCGUCUACGGCAUCCUCGAAAUUG---------UCGUCCUCA---------UCGUGCUCAUCGCUGCCAUUUCCCU ..((((......(((((..((.(((..((((....(((.((....))))).....)))...)---------..)))..))---------..)))))......))))........ ( -20.70, z-score = -1.67, R) >droSec1.super_1 10978314 96 - 14215200 CCGGCAUAAGCCAGCACUCUGUGACCUCUUCAAAUUGCUCGUCUACGGCAUCCUCGAAAUUG---------UCGUCAUCA---------UCGUGCUCAUCGCUGCAAUUUCCCU ..(((....)))(((((..((((((..((((....(((.((....))))).....)))...)---------..)))).))---------..))))).................. ( -20.20, z-score = -1.28, R) >droYak2.chr2R 5432449 96 + 21139217 CCGGCAGAAGCCAGCACUCUGUGACCUCUUCAAACUGCUCGUCUACAGCAUCCUCGAAAUUAUCGUCGUCAUCGUGCUCAUCGCUCCCAUUGCCCU------------------ ..(((((.(((.(((((...(((((..........((((.(....)))))....(((.....)))..))))).)))))....)))....)))))..------------------ ( -20.90, z-score = -1.45, R) >droEre2.scaffold_4845 7707429 96 - 22589142 CCGGAAGAAGCCAGCACUCUGUGACCUCUUCAAACUGUUCGUCUACGGCAUCCUCGAAAUUA---------UCAUCGUCA---------UCGUGCUCAUCGCUCCCAUUUCCCU ..(((((.(((.(((((...(((((.........((((......)))).......((.....---------))...))))---------).)))))....)))....))))).. ( -17.20, z-score = -0.57, R) >droAna3.scaffold_13266 9182303 72 + 19884421 CGGGUAGGAGCCAGCACUCUGUGA------CGAAAUCCUCAUUGAUGGCGUCCUCGUACUCAUCGCUCCCGUUUCCCU------------------------------------ .(((..((((((((....))).((------((..(((......)))..))))............))))).....))).------------------------------------ ( -19.80, z-score = -0.65, R) >dp4.chr3 9045881 99 + 19779522 CGGGCAACAGCCAGCACUCUGUAACUUCCUCAAACUGUUCGUCCACAGCAUCCUCAAAGUUCUCA------UCGUCCUCA---------UCGUGCUCGUCGCUGCCAUUGCCCU .((((((((((.(((((..((.(((((.......((((......))))........)))))..))------..(....).---------..)))))....))))...)))))). ( -22.26, z-score = -2.77, R) >droPer1.super_4 1105864 99 + 7162766 CGGGCAACAGCCAGCACUCUGUAACUUCCUCAAACUGUUCGUCCACAGCAUCCUCAAAGUUCUCA------UCGUCCUCA---------UCGUGCUCGUCGCUGCCAUUGCCCU .((((((((((.(((((..((.(((((.......((((......))))........)))))..))------..(....).---------..)))))....))))...)))))). ( -22.26, z-score = -2.77, R) >droMoj3.scaffold_6496 11131145 114 + 26866924 CAGGUAGCAACCAGCACUCGGUUACUUCAUCCAAAUGCUGAUCGAUAGCAUCCUCGAAAUUAUCAUCUUCGUCGUCCUCAUCAUCCUGGUCGUGCUCAUCACUGCCGUUGCCCU ..((((((...(((((...((.........))...)))))...(((((((....((((.........)))).((.((..........)).)))))).)))......)))))).. ( -24.50, z-score = -1.14, R) >droVir3.scaffold_12875 11196562 108 - 20611582 CGGGCAAUAGCCAGCACUCGGUUACUUCAUCCAGUUGCUGAUCGACGGCAUCCUCAAAGUUGUCA------UCAUCUUCUUCGUCUGGGUCGUGCUCAUCGCUGCCGUUGCCCU .((((((..((.(((..(((((.(((......))).)))))..((((((.........)))))).------..............((((.....))))..)))))..)))))). ( -33.70, z-score = -2.05, R) >consensus CCGGCAGAAGCCAGCACUCUGUGACCUCUUCAAACUGCUCGUCUACGGCAUCCUCGAAAUUAUC_______UCGUCCUCA_________UCGUGCUCAUCGCUGCCAUUGCCCU ..(((....)))(((((..((.........))...((((.......)))).........................................))))).................. ( -7.26 = -7.01 + -0.25)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:30:06 2011