
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 13,139,320 – 13,139,411 |
| Length | 91 |
| Max. P | 0.862000 |

| Location | 13,139,320 – 13,139,411 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 86.72 |
| Shannon entropy | 0.22790 |
| G+C content | 0.47063 |
| Mean single sequence MFE | -28.86 |
| Consensus MFE | -22.24 |
| Energy contribution | -23.10 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.848990 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
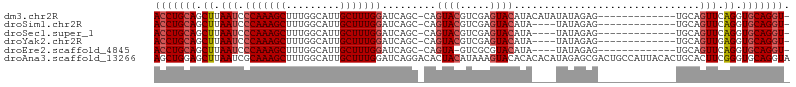
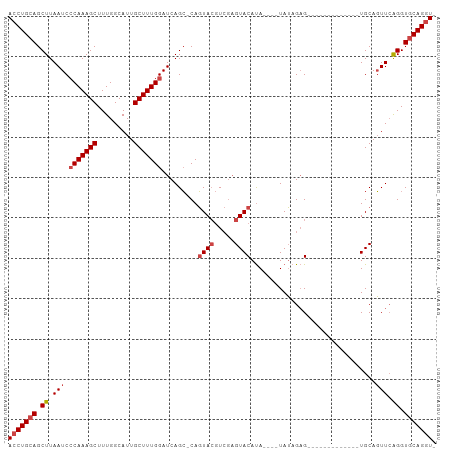
>dm3.chr2R 13139320 91 + 21146708 ACCUGCAGCUUAAUCCCAAAGCUUUGGCAUUGCUUUGGAUCAGC-CAGUACGUCGAGUACAUACAUAUAUAGAG-------------UGCAGUUCAGGUGCAGGU- (((((((.((.(((.(((((((.........)))))))......-..((((.....))))..............-------------....))).)).)))))))- ( -27.60, z-score = -1.74, R) >droSim1.chr2R 11867470 87 + 19596830 ACCUGCAGCUUAAUCCCAAAGCUUUGGCAUUGCUUUGGAUCAGC-CAGUACGUCGAGUACAUA----UAUAGAG-------------UGCAGUUCAGGUGCAGGU- (((((((.((.(((.(((((((.........)))))))......-..((((.....))))...----.......-------------....))).)).)))))))- ( -27.60, z-score = -1.87, R) >droSec1.super_1 10638121 87 + 14215200 ACCUGCAGCUUAAUCCCAAAGCUUUGGCAUUGCUUUGGAUCAGC-CAGUACGUCGAGUACAUA----UAUAGAG-------------UGCAGUUCAGGUGCAGGU- (((((((.((.(((.(((((((.........)))))))......-..((((.....))))...----.......-------------....))).)).)))))))- ( -27.60, z-score = -1.87, R) >droYak2.chr2R 2803392 87 - 21139217 ACCUGCAGCUUAAUCCCAAAGCUUUGGCAUUGCUUUGGAUCAGC-CAGUACGUCGAGUACAUA----UAUAGAG-------------UGCAGUUGAGGUGCAGGU- (((((((.((((((.(((((((.........)))))))......-..((((.....))))...----.......-------------....)))))).)))))))- ( -31.60, z-score = -2.94, R) >droEre2.scaffold_4845 9872631 86 - 22589142 ACCUGCAGCUUAAUCCCAAAGCUUUGGCAUUGCUUUGGAUCAGC-CAGUA-GUCGCGUACAUA----UAUAGAG-------------UGCAGUUCAGGUGCAGGU- (((((((.((.(((.(((((((.........)))))))......-..(((-.((..(((....----))).)).-------------))).))).)).)))))))- ( -27.40, z-score = -1.67, R) >droAna3.scaffold_13266 6398295 106 - 19884421 AGCUGGAGCUUAAUCGCAAAGCUUUGGCAUUGCUUUGGAUCAGGACACUACAUAAAGUACACACACAUAGAGCGACUGCCAUUACACUGCACUUCGGGUGCAGGUA .((..((((((.......))))))..)).((((((((.........(((......))).........)))))))).......(((.((((((.....))))))))) ( -31.37, z-score = -1.96, R) >consensus ACCUGCAGCUUAAUCCCAAAGCUUUGGCAUUGCUUUGGAUCAGC_CAGUACGUCGAGUACAUA____UAUAGAG_____________UGCAGUUCAGGUGCAGGU_ (((((((.((.(((.(((((((.........))))))).........((((.....))))...............................))).)).))))))). (-22.24 = -23.10 + 0.86)

| Location | 13,139,320 – 13,139,411 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 86.72 |
| Shannon entropy | 0.22790 |
| G+C content | 0.47063 |
| Mean single sequence MFE | -26.82 |
| Consensus MFE | -20.97 |
| Energy contribution | -21.80 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.96 |
| SVM RNA-class probability | 0.862000 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
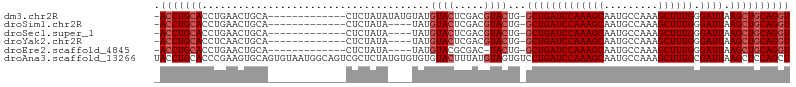
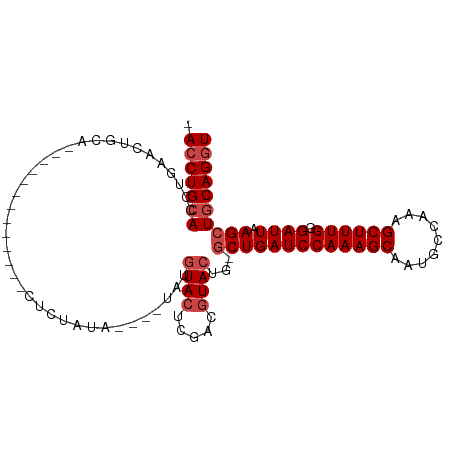
>dm3.chr2R 13139320 91 - 21146708 -ACCUGCACCUGAACUGCA-------------CUCUAUAUAUGUAUGUACUCGACGUACUG-GCUGAUCCAAAGCAAUGCCAAAGCUUUGGGAUUAAGCUGCAGGU -((((((............-------------..........((((((.....)))))).(-(((((((((((((.........))))).))))).)))))))))) ( -28.40, z-score = -2.53, R) >droSim1.chr2R 11867470 87 - 19596830 -ACCUGCACCUGAACUGCA-------------CUCUAUA----UAUGUACUCGACGUACUG-GCUGAUCCAAAGCAAUGCCAAAGCUUUGGGAUUAAGCUGCAGGU -((((((............-------------.......----...((((.....)))).(-(((((((((((((.........))))).))))).)))))))))) ( -26.40, z-score = -2.23, R) >droSec1.super_1 10638121 87 - 14215200 -ACCUGCACCUGAACUGCA-------------CUCUAUA----UAUGUACUCGACGUACUG-GCUGAUCCAAAGCAAUGCCAAAGCUUUGGGAUUAAGCUGCAGGU -((((((............-------------.......----...((((.....)))).(-(((((((((((((.........))))).))))).)))))))))) ( -26.40, z-score = -2.23, R) >droYak2.chr2R 2803392 87 + 21139217 -ACCUGCACCUCAACUGCA-------------CUCUAUA----UAUGUACUCGACGUACUG-GCUGAUCCAAAGCAAUGCCAAAGCUUUGGGAUUAAGCUGCAGGU -((((((............-------------.......----...((((.....)))).(-(((((((((((((.........))))).))))).)))))))))) ( -26.40, z-score = -2.30, R) >droEre2.scaffold_4845 9872631 86 + 22589142 -ACCUGCACCUGAACUGCA-------------CUCUAUA----UAUGUACGCGAC-UACUG-GCUGAUCCAAAGCAAUGCCAAAGCUUUGGGAUUAAGCUGCAGGU -((((((........(((.-------------..(....----...)...)))..-....(-(((((((((((((.........))))).))))).)))))))))) ( -23.70, z-score = -1.30, R) >droAna3.scaffold_13266 6398295 106 + 19884421 UACCUGCACCCGAAGUGCAGUGUAAUGGCAGUCGCUCUAUGUGUGUGUACUUUAUGUAGUGUCCUGAUCCAAAGCAAUGCCAAAGCUUUGCGAUUAAGCUCCAGCU (((((((((.....)))))).))).(((.(((.((.(((((((.........))))))).))..(((((((((((.........)))))).))))).))))))... ( -29.60, z-score = -1.42, R) >consensus _ACCUGCACCUGAACUGCA_____________CUCUAUA____UAUGUACUCGACGUACUG_GCUGAUCCAAAGCAAUGCCAAAGCUUUGGGAUUAAGCUGCAGGU .(((((((......................................((((.....))))...(((((((((((((.........)))))).)))).)))))))))) (-20.97 = -21.80 + 0.83)
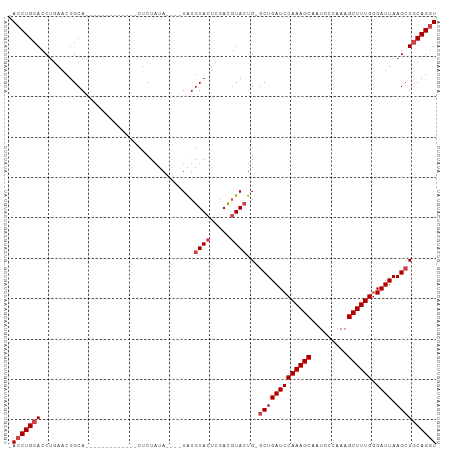
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:29:27 2011