
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,806,035 – 12,806,146 |
| Length | 111 |
| Max. P | 0.989619 |

| Location | 12,806,035 – 12,806,146 |
|---|---|
| Length | 111 |
| Sequences | 8 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 70.23 |
| Shannon entropy | 0.57815 |
| G+C content | 0.43939 |
| Mean single sequence MFE | -29.82 |
| Consensus MFE | -16.71 |
| Energy contribution | -15.91 |
| Covariance contribution | -0.80 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.38 |
| SVM RNA-class probability | 0.989619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 12806035 111 + 21146708 ACUUAUUGCAGUCACUGUUCUGCCCUUA-CAGUAGUGACAACAGGGAUAUUAUCCUUUAGGCUACUCAAUGUUGACUCAUUGGGACGUAAUCGGGC------AAAAGGACAAUUGACG- ..........((((.(((((((((((((-(((((((......((((((...))))))...))))))(((((......)))))....))))..))))------...))))))..)))).- ( -30.70, z-score = -1.86, R) >droSim1.chr2R 11542338 107 + 19596830 ACUUAUUGCAGUCACUGUCCCGCCCUUA-CAGUA----CGGUAAGGAUAUUAUCCUUUAGGCUACUCAAUGUUGACUCAUUGAGACGUAAUCGGGC------AAAAGGACAAUUGACG- ..........((((.(((((.(((((((-((((.----....((((((...))))))...))).(((((((......)))))))..))))..))))------....)))))..)))).- ( -34.30, z-score = -3.53, R) >droSec1.super_1 10310036 111 + 14215200 ACUUAUUGCAGUCACUGUCCUGCCCUUA-CAGUAGUCACUGUAAGGAUAUUAUCCUUUAGGCUACUCAAUGUUGACUCAUUGAGACGUAAUCGGGC------AAAAGGACAAUUGACG- ..........((((.(((((((((((((-(.((((((.....((((((...))))))..))))))((((((......))))))...))))..))))------...))))))..)))).- ( -37.80, z-score = -4.31, R) >droYak2.chr2R 2474953 99 - 21139217 ACUUAUUGCAGUCACUGUUGUGUCCUUA-C------------GAGGAUAUAAUCCUUAAGACUACUCAAUGUUGACUCAUUGAGACGUAAUCGGGC------AAAACGACAAUUGACG- ..........((((.(((((((((((((-(------------((((((...)))))........(((((((......))))))).)))))..))))------...))))))..)))).- ( -29.30, z-score = -3.46, R) >droEre2.scaffold_4845 9550088 110 - 22589142 ACUUAUUGCAGUCACUGUUCUGCCCUU--CAGUAAUGACAGCGAGGAUAUUAUCCUCAAGGCUACUGAAUGUGGACUCAUUGAGACGUAAUCGGGC------AAAAGGACAAUUGACG- ..........((((.((((((((((.(--((....))).(((((((((...))))))...))).((.((((......)))).))........))))------...))))))..)))).- ( -30.80, z-score = -1.60, R) >droAna3.scaffold_13266 6093276 111 - 19884421 ACUUACUGCAGUCACUGUACCGGCUUUGGUCACAAUGACAAUGAUGAUAAUAUCGUUAAUGGUACUCAGUGCUGACUCAAUGAGACGUAAUCGGCC------AA--GGACAUACGACAU ..........(((...(((....((((((((.......((.(((((((...))))))).))((.((((.((......)).))))))......))))------))--))...)))))).. ( -24.10, z-score = 0.37, R) >dp4.chr3 6754079 113 + 19779522 GCCUGUUGUAGUCAGUGGCUGGAUUACGCUAAUGACAUU--UGAUGAUAACAUCGCUU---CUACUCAGCAAGGACUCAUUGAGACAUAAUCGGCCGCCCACAAAAGGACACCCGAUG- .(((.((((.(...(((((((.((((.((((((((....--.((((....))))(((.---......)))......))))).)).).)))))))))))).)))).)))..........- ( -26.50, z-score = -0.20, R) >droPer1.super_2 6976483 113 + 9036312 GCCUGUUUUAGUCAGUGGCUGGAUUACGCUAAUGACAUU--UGAUGAUAACAUAGCUU---CUACUCAGCAAGGACUCAUUGAGACAUAAUCGGCCGCCCACAAAAGGACACCCGAUG- ...(((((((((.(((.((((..(...((((.((..(((--....)))..))))))..---..)..))))....))).)))))))))..(((((..(.((......)).)..))))).- ( -25.10, z-score = -0.31, R) >consensus ACUUAUUGCAGUCACUGUCCUGCCCUUA_CAGUAAUGAC__UGAGGAUAUUAUCCUUUAGGCUACUCAAUGUUGACUCAUUGAGACGUAAUCGGGC______AAAAGGACAAUUGACG_ ..........(((..(((((.((((.................((((((...)))))).......(((((((......)))))))........))))..........)))))...))).. (-16.71 = -15.91 + -0.80)

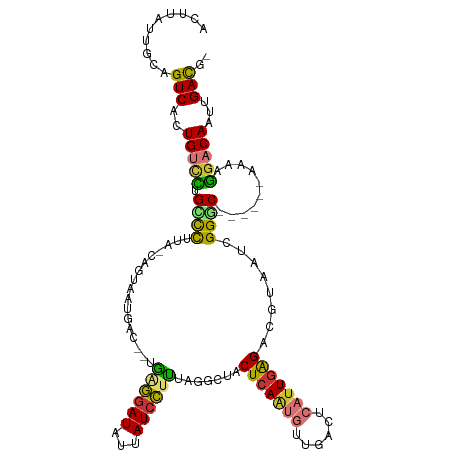
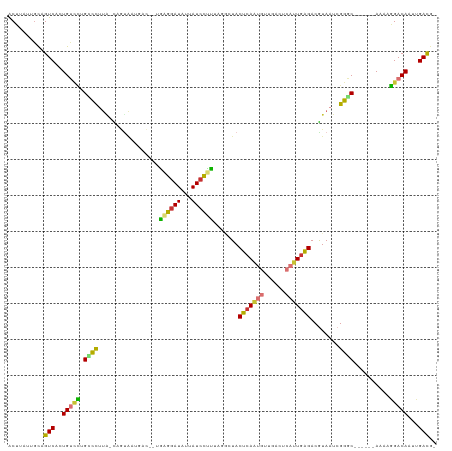
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:28:29 2011