| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,778,766 – 12,778,908 |
| Length | 142 |
| Max. P | 0.989405 |

| Location | 12,778,766 – 12,778,908 |
|---|---|
| Length | 142 |
| Sequences | 6 |
| Columns | 149 |
| Reading direction | reverse |
| Mean pairwise identity | 80.92 |
| Shannon entropy | 0.36118 |
| G+C content | 0.40801 |
| Mean single sequence MFE | -34.76 |
| Consensus MFE | -23.28 |
| Energy contribution | -25.70 |
| Covariance contribution | 2.42 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.989405 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
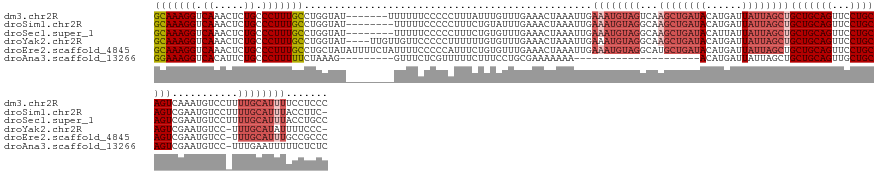
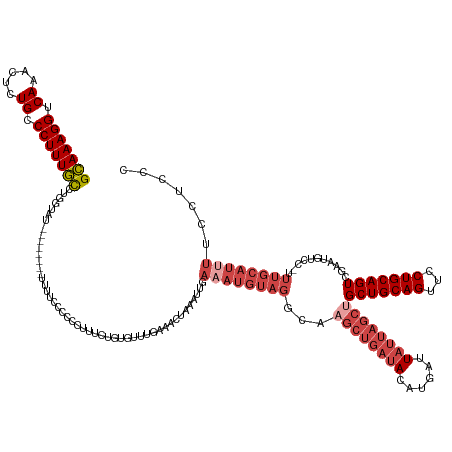
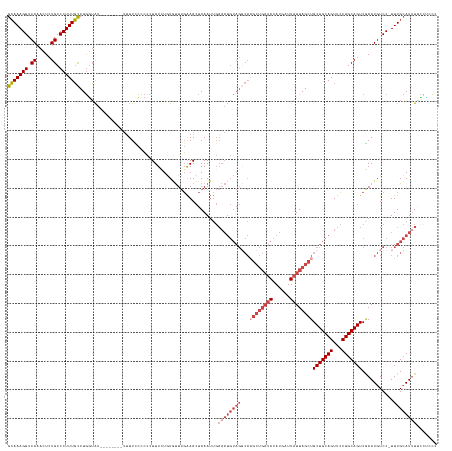
>dm3.chr2R 12778766 142 - 21146708 GCAAAGGUCAAACUCUGCCCUUUGCCUGGUAU-------UUUUUUCCCCCUUUAUUUGUUUGAAACUAAAUUGAAAUGUAGUCAAGCUGAUACAUGAUUAUUAGCUGCUGCAGUUCCUGCAGUCAAAUGUCCUUUUGCAUUUUCCUCCC (((((((.((.....)).))))))).......-------.................................(((((((((...((((((((......))))))))(((((((...)))))))...........)))))))))...... ( -32.00, z-score = -2.56, R) >droSim1.chr2R 11514417 140 - 19596830 GCAAAGGUCAAACUCUGCCCUUUGCCUGGUAU--------UUUUUCCCCCUUUCUGUAUUUGAAACUAAAUUGAAAUGUAGGCAAGCUGAUACAUGAUUAUUAGCUGCUGCAGUUCCUGCAGUCGAAUGUCCUUUUGCAUUUACCUUC- (((((((.((.....)).)))))))..((((.--------.............((((((((.((......)).))))))))(((((((((((......))))))))(((((((...)))))))............)))...))))...- ( -36.20, z-score = -2.90, R) >droSec1.super_1 10282899 141 - 14215200 GCAAAGGUCAAACUCUGCCCUUUGCCUGGUAU--------UUUUUCCCCCUUUCUGUGUUUGAAACUAAAUUGAAAUGUAGGCAAGCUGAUACAUUAUUAUUAGCUGCUGCAGUUCCUGCAGUCGAAUGUCCUUUUGCAUUUACCUGCC (((((((.((.....)).)))))))..((((.--------.............(((..(((.((......)).)))..)))(((((((((((......))))))))(((((((...)))))))............)))...)))).... ( -37.50, z-score = -2.98, R) >droYak2.chr2R 4943279 143 + 21139217 GCAAAGGUCAAACUCUGCCCUUUGCCUGGUAU----UUGUUGUUCCCCCUUUUUUGUGUUUGAAACUAAAUUGAAAUGUAGGCAAGCUGAUACAUGAUUAUUAGCUGCUGCAGUUCCUGCAGUCGAAUGUCC-UUUGCAUAUUUUCCC- (((((((.((............(((((((.((----.....)).)).........(..(((.((......)).)))..))))))((((((((......))))))))(((((((...)))))))....)).))-)))))..........- ( -36.10, z-score = -2.38, R) >droEre2.scaffold_4845 9520363 148 + 22589142 GCAAAGGUCAAACUCUGCCCUUUGCCUGCUAUAUUUUCUAUUUUCCCCCAUUUCUGUGUUUGAAACUAAAUUGAAAUGUAGGCAUGCUGAUACAUGAUUAUUAGCUGCUGCAGUUCCUGCAGUCGAAUGUCC-UUUGCAUUUGCCGCCC (((((((.((............(((((((............((((...(((....)))...))))............))))))).(((((((......))))))).(((((((...)))))))....)).))-)))))........... ( -39.55, z-score = -3.34, R) >droAna3.scaffold_13266 6064802 118 + 19884421 GGAAAGGUCACAUUCUGCCCUUUUUCUAAAG---------GUUUCUCGUUUUUCUUUCCUGCGAAAAAAA---------------------ACAUGAUUAUUAGCUGCUGCAGUUGCUGCAGUCGAAUGUCC-UUUGAAUUUUUCUCUC ..(((((..((((((.(((((((....))))---------)......((((((.((((....))))))))---------------------))..........)).(((((((...))))))).))))))))-)))............. ( -27.20, z-score = -1.62, R) >consensus GCAAAGGUCAAACUCUGCCCUUUGCCUGGUAU________UUUUCCCCCCUUUCUGUGUUUGAAACUAAAUUGAAAUGUAGGCAAGCUGAUACAUGAUUAUUAGCUGCUGCAGUUCCUGCAGUCGAAUGUCC_UUUGCAUUUUCCUCCC (((((((.((.....)).)))))))................................................((((((((...((((((((......))))))))(((((((...)))))))...........))))))))....... (-23.28 = -25.70 + 2.42)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:28:24 2011