
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,743,760 – 12,743,817 |
| Length | 57 |
| Max. P | 0.623065 |

| Location | 12,743,760 – 12,743,817 |
|---|---|
| Length | 57 |
| Sequences | 12 |
| Columns | 58 |
| Reading direction | reverse |
| Mean pairwise identity | 85.15 |
| Shannon entropy | 0.31131 |
| G+C content | 0.34360 |
| Mean single sequence MFE | -5.02 |
| Consensus MFE | -4.14 |
| Energy contribution | -4.23 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.623065 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 12743760 57 - 21146708 UUUACAUACCAUAUUUCUAAUGGCUUUUCU-GUUUUUCCGUCAGCCGACACACAUUCA ........((((.......)))).......-........(((....)))......... ( -4.50, z-score = -0.72, R) >droEre2.scaffold_4845 9480866 57 + 22589142 UUUACAUACCAUAUUUCUAAUGGCUUUUCU-GUUUUUCCGUCAGCCGACACACAUUCA ........((((.......)))).......-........(((....)))......... ( -4.50, z-score = -0.72, R) >droYak2.chr2R 4906968 57 + 21139217 UUUACGUACCAUAUUUCUAAUGGCUUUUCU-GUUUUUCCGUCAGCCGACACACAUUCA .....((.((((.......)))).......-........(((....)))))....... ( -4.60, z-score = -0.28, R) >droSec1.super_1 10248153 57 - 14215200 UUUACAUACCAUAUUUCUAAUGGCUCUUCU-GUUUUUCCGUCAGCCGACAAACAUUCA ........((((.......))))......(-((((.((.(....).)).))))).... ( -5.20, z-score = -0.96, R) >droSim1.chr2R 11479052 57 - 19596830 UUUACAUACCAUAUUUCUAAUGGCUUUUCU-GUUUUUCCGUCAGCCGACACACAUUCA ........((((.......)))).......-........(((....)))......... ( -4.50, z-score = -0.72, R) >dp4.Unknown_singleton_1898 54496 57 + 80150 UCUACGUACCAUAUUUUUAAUGGUUUUUAU-AUUUUUCCGUCAGCCGACACACAUUCA .....(((((((.......)))))......-........(((....)))))....... ( -5.70, z-score = -1.52, R) >droPer1.super_4 6567781 57 - 7162766 UCUACGUACCAUAUUUUUAAUGGUUUUUAU-AUUUUUCCGUCAGCCGACACACAUUCA .....(((((((.......)))))......-........(((....)))))....... ( -5.70, z-score = -1.52, R) >droAna3.scaffold_13266 6029057 57 + 19884421 UUUACAUACCAUAUUUCUAAUGGUUUUUCU-GUUUUUUCUUCAGCCGACACACAUUCA .......(((((.......)))))......-........................... ( -3.10, z-score = -0.45, R) >droWil1.scaffold_180700 6418816 58 - 6630534 UUUACAUACCAUAUUUUCAAUGGUUUUUAUCUUUUUUUCGUCAGCCGACACACAUUCA .......(((((.......)))))...............(((....)))......... ( -5.60, z-score = -2.65, R) >droVir3.scaffold_12875 9967693 56 + 20611582 UUUACAUACCAUAUUUUCAAUGGUUUUAUCUUUUUUUUCGUCAGCCGACAUGCUUU-- .......(((((.......)))))...............(((....))).......-- ( -5.60, z-score = -1.61, R) >droMoj3.scaffold_6496 16317199 58 - 26866924 UUUACGUACCAUAUUUUCAAUGGUUUUAUCUUUUUUUUCGUCAGCCGACAUGCUUUUG .......(((((.......)))))...............(((....)))......... ( -5.60, z-score = -1.01, R) >droGri2.scaffold_15245 14568472 53 + 18325388 UUUACGUACCAUAUUUUCAAUGGUUUUAUCUUUUUUUUCGUCAGCCGACAUGC----- .......(((((.......)))))...............(((....)))....----- ( -5.60, z-score = -1.07, R) >consensus UUUACAUACCAUAUUUCUAAUGGUUUUUCU_GUUUUUCCGUCAGCCGACACACAUUCA ........((((.......))))................(((....)))......... ( -4.14 = -4.23 + 0.08)


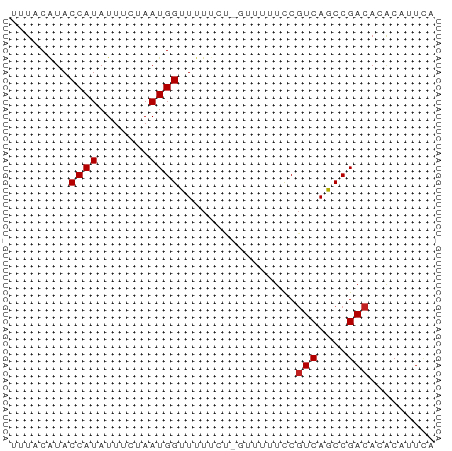
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:28:19 2011