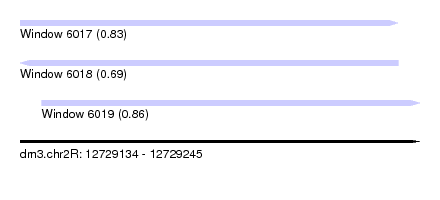

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,729,134 – 12,729,245 |
| Length | 111 |
| Max. P | 0.858456 |

| Location | 12,729,134 – 12,729,239 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 86.71 |
| Shannon entropy | 0.17491 |
| G+C content | 0.42002 |
| Mean single sequence MFE | -17.12 |
| Consensus MFE | -16.19 |
| Energy contribution | -15.75 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.834122 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
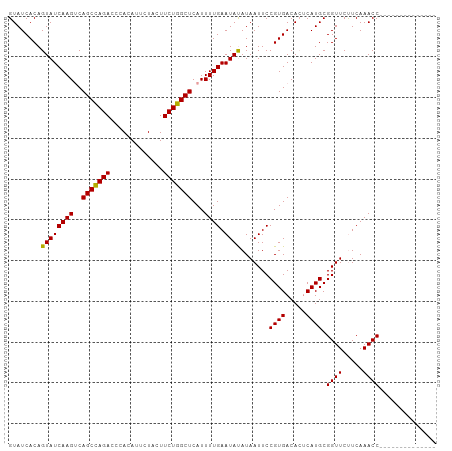
>dm3.chr2R 12729134 105 + 21146708 GUAUCACAGUAUCAAGUCAGCUAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUAUAUAAUUCCGUGACAGUCAUGCGGUUCUUCAAACCAAAAAAGUACAGAU ((((.......(((((..(((((((.............)))))))...)))))...........((((.....)))).((((.....))))......)))).... ( -16.72, z-score = -0.64, R) >droSim1.chr2R 11464505 91 + 19596830 GUAUCACAGUAUCAAGUCAGCCAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUAUAUAAUUCCGUGACACUCAUGCGGUUCCUCAAACC-------------- ((.((((......((((.(((((((.............))))))).)))).((((.....)))).)))).))......((((.....))))-------------- ( -17.12, z-score = -1.96, R) >droSec1.super_1 10233708 91 + 14215200 GUAUCACAGUAUCAAGGCAGCCAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUACAUAAUUCCGUGACACGCAUGCGGUUCUUCAAACC-------------- ........(((((((((.(((((((.............)))))))..))))).))))......((((.........))))...........-------------- ( -17.52, z-score = -1.25, R) >consensus GUAUCACAGUAUCAAGUCAGCCAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUAUAUAAUUCCGUGACACUCAUGCGGUUCUUCAAACC______________ ........((((((((..(((((((.............)))))))...)))).)))).......((((.....)))).((((.....)))).............. (-16.19 = -15.75 + -0.44)


| Location | 12,729,134 – 12,729,239 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 86.71 |
| Shannon entropy | 0.17491 |
| G+C content | 0.42002 |
| Mean single sequence MFE | -23.61 |
| Consensus MFE | -21.40 |
| Energy contribution | -22.07 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.685411 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
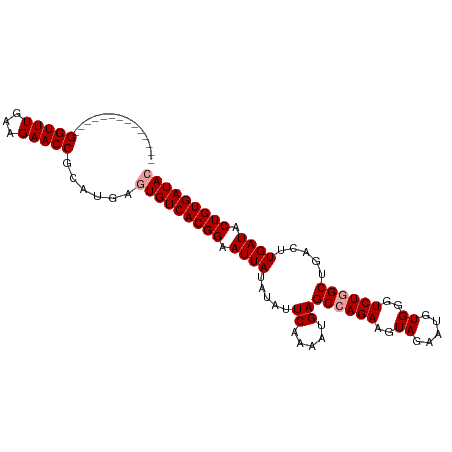
>dm3.chr2R 12729134 105 - 21146708 AUCUGUACUUUUUUGGUUUGAAGAACCGCAUGACUGUCACGGAAUUAUAUAUUCAAAAUGAGCCAGAAGUAGAAUGUGGGUCUAGCUGACUUGAUACUGUGAUAC ..............(((((...))))).......((((((((.(((((((((((.................)))))))((((.....)))))))).)))))))). ( -19.83, z-score = 0.35, R) >droSim1.chr2R 11464505 91 - 19596830 --------------GGUUUGAGGAACCGCAUGAGUGUCACGGAAUUAUAUAUUCAAAAUGAGCCAGAAGUAGAAUGUGGGUCUGGCUGACUUGAUACUGUGAUAC --------------(((((...)))))......(((((((((.((((.....((.....))((((((..((.....))..)))))).....)))).))))))))) ( -24.80, z-score = -2.12, R) >droSec1.super_1 10233708 91 - 14215200 --------------GGUUUGAAGAACCGCAUGCGUGUCACGGAAUUAUGUAUUCAAAAUGAGCCAGAAGUAGAAUGUGGGUCUGGCUGCCUUGAUACUGUGAUAC --------------(((((...)))))......(((((((((.((((.(((.((.....))((((((..((.....))..)))))))))..)))).))))))))) ( -26.20, z-score = -1.97, R) >consensus ______________GGUUUGAAGAACCGCAUGAGUGUCACGGAAUUAUAUAUUCAAAAUGAGCCAGAAGUAGAAUGUGGGUCUGGCUGACUUGAUACUGUGAUAC ..............(((((...)))))......(((((((((.((((.....((.....))((((((..((.....))..)))))).....)))).))))))))) (-21.40 = -22.07 + 0.67)

| Location | 12,729,140 – 12,729,245 |
|---|---|
| Length | 105 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 82.37 |
| Shannon entropy | 0.22739 |
| G+C content | 0.42185 |
| Mean single sequence MFE | -17.25 |
| Consensus MFE | -16.19 |
| Energy contribution | -15.75 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.27 |
| Structure conservation index | 0.94 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.94 |
| SVM RNA-class probability | 0.858456 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
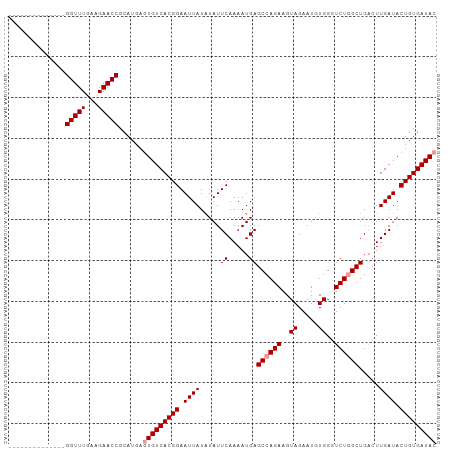
>dm3.chr2R 12729140 105 + 21146708 CAGUAUCAAGUCAGCUAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUAUAUAAUUCCGUGACAGUCAUGCGGUUCUUCAAACCAAAAAAGUACAGAUUGAUAU ..(((((((((((((((((.............)))))).......((((.....))))..))))........((((.....)))).............))))))) ( -18.42, z-score = -0.80, R) >droSim1.chr2R 11464511 85 + 19596830 CAGUAUCAAGUCAGCCAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUAUAUAAUUCCGUGACACUCAUGCGGUUCCUCAAACC-------------------- .....(((((..(((((((.............)))))))...)))))......(((..((((.....))))..))).........-------------------- ( -15.82, z-score = -1.62, R) >droSec1.super_1 10233714 85 + 14215200 CAGUAUCAAGGCAGCCAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUACAUAAUUCCGUGACACGCAUGCGGUUCUUCAAACC-------------------- ..(((((((((.(((((((.............)))))))..))))).))))......((((.........))))...........-------------------- ( -17.52, z-score = -1.40, R) >consensus CAGUAUCAAGUCAGCCAGACCCACAUUCUACUUCUGGCUCAUUUUGAAUAUAUAAUUCCGUGACACUCAUGCGGUUCUUCAAACC____________________ ..((((((((..(((((((.............)))))))...)))).)))).......((((.....)))).((((.....)))).................... (-16.19 = -15.75 + -0.44)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:28:17 2011