
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,630,563 – 12,630,665 |
| Length | 102 |
| Max. P | 0.676586 |

| Location | 12,630,563 – 12,630,665 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 69.65 |
| Shannon entropy | 0.54762 |
| G+C content | 0.37178 |
| Mean single sequence MFE | -23.90 |
| Consensus MFE | -8.69 |
| Energy contribution | -7.44 |
| Covariance contribution | -1.25 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.676586 |
| Prediction | RNA |
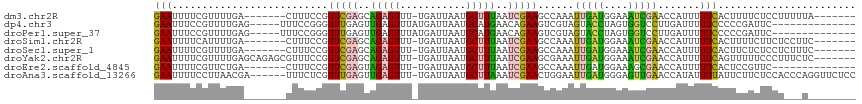
Download alignment: ClustalW | MAF
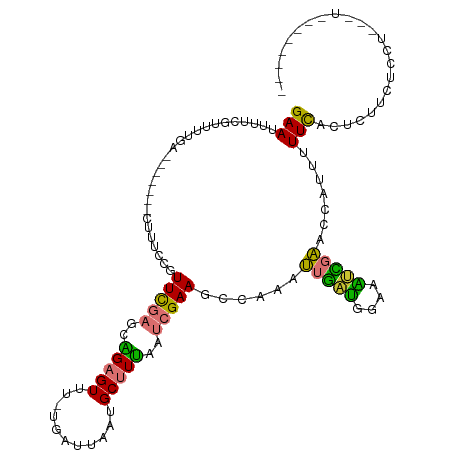
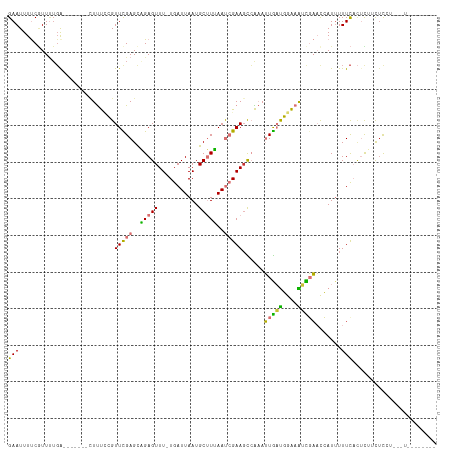
>dm3.chr2R 12630563 102 - 21146708 GAAUUUUCGUUUUGA-------CUUUCCGUUCGAGCAGAGUUU-UGAUUAAUGCUUUAAUCGAAGCCAAAUUGAUGGAAAUCGAACCAUUUUUCACUUUUCUCCUUUUUA------- (((.......(((((-------.(((((((.(((...(.((((-(((((((....))))))))))))...)))))))))))))))......)))................------- ( -22.62, z-score = -2.23, R) >dp4.chr3 695443 98 + 19779522 GAAUUUCCGUUUUGAG-----UUUCCGGGUUUGAGUUGAGUUUAUGAUUAAUGCAUGAACAGAAGUCGUAGUACCUAGUGGUCCUUGAUUUUUCCCCCGAUUC-------------- (((.(((......)))-----.)))((((..........(((((((.......)))))))((((((((.((.(((....))).))))))))))..))))....-------------- ( -22.30, z-score = -1.63, R) >droPer1.super_37 673521 98 + 760358 GAAUUUCCGUUUUGAG-----UUUCCGGGUUUGAGUUGAGUUUAUGAUUAAUGCAUGAACAGAAGUCGUAGUACCUAGUGGUCCUUGAUUUUUCCCCCGAUUC-------------- (((.(((......)))-----.)))((((..........(((((((.......)))))))((((((((.((.(((....))).))))))))))..))))....-------------- ( -22.30, z-score = -1.63, R) >droSim1.chr2R 11365346 102 - 19596830 GAAUUUUCAUUUUGA-------CUUUCCGUUCGAGCAGAGUUU-UGAUUAAUGCUUUAAUCGAAGCCAAAUUGAUGGAAAUCGAACCAUUUUUCACUUUUCUUCUCCUUC------- (((.......(((((-------.(((((((.(((...(.((((-(((((((....))))))))))))...)))))))))))))))......)))................------- ( -22.62, z-score = -2.22, R) >droSec1.super_1 10136027 102 - 14215200 GAAUUUUCGUUUUGA-------CUUUCCGUUCGAGCAGAGUUU-UGAUUAAUGCUUUAAUCGAAGCCAAAUUGAUGGAAAUCGAACCAUUUUUCACUUCUCUCCUCUUUC------- (((.......(((((-------.(((((((.(((...(.((((-(((((((....))))))))))))...)))))))))))))))......)))................------- ( -22.62, z-score = -1.78, R) >droYak2.chr2R 4791459 109 + 21139217 GAAUUUUCGUUUUGAGCAGAGCGUUUCCGUUCGAGCAGAGUUU-UGAUUAAUGCUUUAAUCGAAGCGAAAUUGAUGGAAAUCGAACCAUUUUUCAGUUUUUCCCUUUCUC------- (((.(((((((((((..((((((((.....(((((......))-)))..))))))))..)))))))))))(((((....))))).......)))................------- ( -27.20, z-score = -1.47, R) >droEre2.scaffold_4845 9367809 95 + 22589142 GAAUUUUCGUUCUGA-------CUUUCCGUUCGAGUAGAGUUU-UGAUUAAUGCUUUAAUCGAAGCCAAAUUGAUGGAAAGCGAACCAUUUUUCACUCCGUUC-------------- (((.....((((.(.-------((((((((.(((...(.((((-(((((((....))))))))))))...)))))))))))))))).....))).........-------------- ( -26.60, z-score = -3.12, R) >droAna3.scaffold_13266 5920806 110 + 19884421 GAAUUUUCCUUAACGA------UUUCUCGUUUGAGUUGAGUUU-UGAUUAAUGCUUAAAUCGAACUGGAAUUGAUGGGAGUUGAACCAUAUUUUAUUCUUCUCCACCCAGGUUCUCC .....(((((...(((------((((..((((((.((((((..-........)))))).)))))).)))))))..)))))..(((((......................)))))... ( -24.95, z-score = -1.26, R) >consensus GAAUUUUCGUUUUGA_______CUUUCCGUUCGAGCAGAGUUU_UGAUUAAUGCUUUAAUCGAAGCCAAAUUGAUGGAAAUCGAACCAUUUUUCACUCUUCUCCU___U________ (((..........................(((((..(((((...........)))))..)))))......(((((....))))).......)))....................... ( -8.69 = -7.44 + -1.25)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:28:03 2011