
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,619,645 – 12,619,746 |
| Length | 101 |
| Max. P | 0.732864 |

| Location | 12,619,645 – 12,619,745 |
|---|---|
| Length | 100 |
| Sequences | 7 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 72.59 |
| Shannon entropy | 0.49926 |
| G+C content | 0.50580 |
| Mean single sequence MFE | -25.85 |
| Consensus MFE | -15.58 |
| Energy contribution | -15.79 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.11 |
| Mean z-score | -0.89 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.520600 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
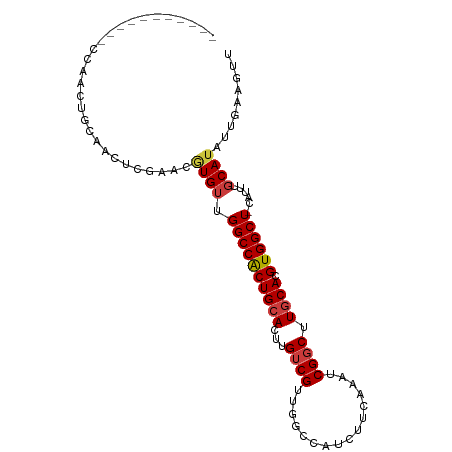
>dm3.chr2R 12619645 100 + 21146708 CAACUGCAACUCGAACUGCAAGUCUAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU-CAUUUGCAUAUUGAAGUU .((((.(((.....((((((((((.....((.(((((((.((....)).).))))))....))....)))))))).)).((.-.....))...))).)))) ( -27.10, z-score = -0.75, R) >droAna3.scaffold_13266 5907284 86 - 19884421 -----------CAACAUCAGACUCGAACCUGUUGGCCACUGCACUUGUCGUUUUAGCCAACU----CGCCAUGCACGUGGCUAUAGUUUCAUACAAGUGCC -----------...((.(((........))).))......((((((((.....((((((...----.((...))...)))))).........)))))))). ( -20.84, z-score = -0.71, R) >droEre2.scaffold_4845 9356073 100 - 22589142 CAACGGCAACUCGAACUGCAAGCCGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU-CAUUUGCAUAUUGAAAUU (((..((((.....((((((((((((...((.(((((((.((....)).).))))))....))..)))))))))).))....-...))))...)))..... ( -34.30, z-score = -2.59, R) >droYak2.chr2R 4779434 100 - 21139217 CGACGGCAACUCGAACUGCAAGUUGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU-CAUUUGCAUAUUGAAGUU (((.(....)))).((((((((((((...((.(((((((.((....)).).))))))....))..)))))))))).)).((.-.....))........... ( -30.70, z-score = -0.92, R) >droSec1.super_1 10125499 88 + 14215200 ------------CAACUGCAACUCGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUUCACGUGGCU-CAUUUGCAUAUUGAAGUU ------------.((((.(((...........(((((((.((....)).).))))))....(((((.((((.......))))-.)))))....))).)))) ( -19.40, z-score = 0.31, R) >droPer1.super_37 660474 84 - 760358 ------------CAACUCGAACACGCACAUGUUGGCCGCUGCACUUGUCGCUCGAACUCCUC----CGGCAUGCACGUGGCU-CAUUUGCAUUUUGUGGUU ------------.((((((((...(((.(((..((((((((((..(((((...(....)...----))))))))).))))))-))).)))..)))).)))) ( -24.80, z-score = -0.74, R) >dp4.chr3 680632 84 - 19779522 ------------CAACUCGAACACGCACAUGUUGGCCGCUGCACUUGUCGCUCGAACUCCUC----CGGCAUGCACGUGGCU-CAUUUGCAUUUUGUUGUU ------------.(((.((((...(((.(((..((((((((((..(((((...(....)...----))))))))).))))))-))).)))..))))..))) ( -23.80, z-score = -0.81, R) >consensus ___________CCAACUGCAACUCGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU_CAUUUGCAUAUUGAAGUU ............................((((.((((((((((...((((................)))).)))).))))))......))))......... (-15.58 = -15.79 + 0.21)


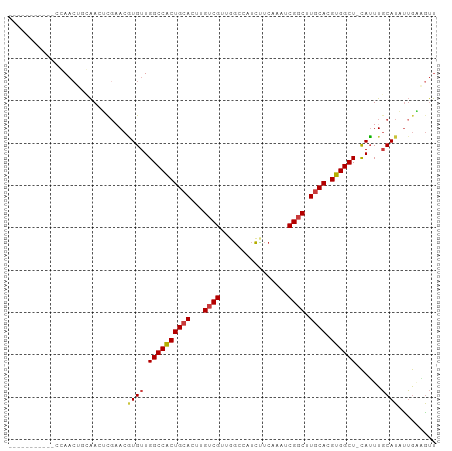
| Location | 12,619,646 – 12,619,746 |
|---|---|
| Length | 100 |
| Sequences | 7 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 72.89 |
| Shannon entropy | 0.49536 |
| G+C content | 0.50414 |
| Mean single sequence MFE | -26.78 |
| Consensus MFE | -15.58 |
| Energy contribution | -15.79 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.732864 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 12619646 100 + 21146708 AACUGCAACUCGAACUGCAAGUCUAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU-CAUUUGCAUAUUGAAGUUG .....((((((((((((((((((.....((.(((((((.((....)).).))))))....))....)))))))).)).((.-.....))...))).))))) ( -29.30, z-score = -1.51, R) >droAna3.scaffold_13266 5907285 86 - 19884421 -----------AACAUCAGACUCGAACCUGUUGGCCACUGCACUUGUCGUUUUAGCCAACU----CGCCAUGCACGUGGCUAUAGUUUCAUACAAGUGCCA -----------..((.(((........))).))......((((((((.....((((((...----.((...))...)))))).........)))))))).. ( -20.84, z-score = -0.65, R) >droEre2.scaffold_4845 9356074 100 - 22589142 AACGGCAACUCGAACUGCAAGCCGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU-CAUUUGCAUAUUGAAAUUG ...(....)((((((((((((((((...((.(((((((.((....)).).))))))....))..)))))))))).)).((.-.....))...))))..... ( -34.20, z-score = -2.58, R) >droYak2.chr2R 4779435 100 - 21139217 GACGGCAACUCGAACUGCAAGUUGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU-CAUUUGCAUAUUGAAGUUG .....((((((((((((((((((((...((.(((((((.((....)).).))))))....))..)))))))))).)).((.-.....))...))).))))) ( -32.00, z-score = -1.42, R) >droSec1.super_1 10125500 88 + 14215200 ------------AACUGCAACUCGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUUCACGUGGCU-CAUUUGCAUAUUGAAGUUG ------------.....(((((..((..((.(((((((.((....)).).))))))....(((((.((((.......))))-.)))))))..))..))))) ( -21.70, z-score = -0.50, R) >droPer1.super_37 660475 84 - 760358 ------------AACUCGAACACGCACAUGUUGGCCGCUGCACUUGUCGCUCGAACUCCUC----CGGCAUGCACGUGGCU-CAUUUGCAUUUUGUGGUUG ------------((((((((...(((.(((..((((((((((..(((((...(....)...----))))))))).))))))-))).)))..)))).)))). ( -25.20, z-score = -0.90, R) >dp4.chr3 680633 84 - 19779522 ------------AACUCGAACACGCACAUGUUGGCCGCUGCACUUGUCGCUCGAACUCCUC----CGGCAUGCACGUGGCU-CAUUUGCAUUUUGUUGUUG ------------(((.((((...(((.(((..((((((((((..(((((...(....)...----))))))))).))))))-))).)))..))))..))). ( -24.20, z-score = -0.97, R) >consensus ____________AACUGCAACUCGAACGUGUUGGCCACUGCACUUGUCGUUGGCCAUCUUCAAAUCGGCUUGCACGUGGCU_CAUUUGCAUAUUGAAGUUG ...........................((((.((((((((((...((((................)))).)))).))))))......)))).......... (-15.58 = -15.79 + 0.21)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:28:00 2011