
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,540,961 – 12,541,017 |
| Length | 56 |
| Max. P | 0.805425 |

| Location | 12,540,961 – 12,541,017 |
|---|---|
| Length | 56 |
| Sequences | 11 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 68.46 |
| Shannon entropy | 0.69035 |
| G+C content | 0.39494 |
| Mean single sequence MFE | -9.51 |
| Consensus MFE | -5.70 |
| Energy contribution | -5.55 |
| Covariance contribution | -0.15 |
| Combinations/Pair | 1.45 |
| Mean z-score | -0.35 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.805425 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 12540961 56 - 21146708 ACGUGACUUGCUUUCCAAUAUGGUUACUUUGGUGUGUAAGUUGAUUAAAAUACUCC- ....(((((((...((((...(....).))))...)))))))..............- ( -10.90, z-score = -1.51, R) >droSim1.chr2R 11274644 56 - 19596830 ACGUGACUUGCUUUCCAAUAUGGUUACUUUGGUGUGUAAGUUGAUCAAAAUACUCC- ....(((((((...((((...(....).))))...)))))))..............- ( -10.90, z-score = -1.24, R) >droSec1.super_1 10039004 56 - 14215200 ACGUGACUUGCUUUCCAAUAUGGUUACUUUGGUGUGUAAGUUGAUCAAAAUACUCC- ....(((((((...((((...(....).))))...)))))))..............- ( -10.90, z-score = -1.24, R) >droYak2.chr2R 4698836 56 + 21139217 ACGUGACUUGCUUUCCAAUAUGGUCACUUUGGUGUGUAAGUUGAUUGAAAUACUCC- ....(((((((...((((...(....).))))...)))))))..............- ( -10.90, z-score = -0.95, R) >droEre2.scaffold_4845 9270069 56 + 22589142 ACGUGACUUGCUUUCCAAUAUGGUCACUUUGGUGUGUAAGUUGAUUGCAACACUCC- ..((((((.............))))))...(((((((((.....)))).)))))..- ( -13.12, z-score = -1.08, R) >dp4.chr3 596456 56 + 19779522 UCAGGACUUGCUCUCUAACAUGGUGACCCUGGUUUGUAAGUUCAUCCAUUUUUCUU- ...((((((((...(((....(....)..)))...)))))))).............- ( -12.00, z-score = -1.12, R) >droPer1.super_37 576098 56 + 760358 UCAGGACUUGCUCUCUAACAUGGUGACCCUGGUUUGUAAGUUCAUCCAUUUUUCUU- ...((((((((...(((....(....)..)))...)))))))).............- ( -12.00, z-score = -1.12, R) >droWil1.scaffold_180569 647871 54 + 1405142 UCAUGAUUUGCUCUCUAAUAUGGUCACUUUGGUCUGUAAGUUCCAUGGUUAAUC--- (((((((((((...((((...(....).))))...))))))..)))))......--- ( -8.20, z-score = -0.60, R) >droVir3.scaffold_12875 17809754 57 + 20611582 CGUCGAUUUGCUCUCAAAUAUGGUCACCCUAGUCUGUAAGUUUGGUUUUGGAUAACU .(((.(...(((......(((((.(......).))))).....)))..).))).... ( -4.40, z-score = 1.95, R) >droMoj3.scaffold_6496 22410370 57 - 26866924 UGUAGAUCUGCUGUCAAAUAUGGUCACCCUAGUCUGUGAGUUGAGCGACCGAUAAUA .(((....))).........(((((.(.(.(.((...)).).).).)))))...... ( -6.30, z-score = 1.54, R) >droGri2.scaffold_15112 3362380 55 - 5172618 CGUGGAUCUAUUAACAAACAUGGUCACCCUAGUCUGUAAGUUGAA--AUUGUCGAUA .((.(((...(((((..(((.((....)).....)))..))))).--...))).)). ( -5.00, z-score = 1.49, R) >consensus ACGUGACUUGCUUUCCAAUAUGGUCACUUUGGUGUGUAAGUUGAUCGAAAUACUCC_ ....(((((((...(((....(....)..)))...)))))))............... ( -5.70 = -5.55 + -0.15)


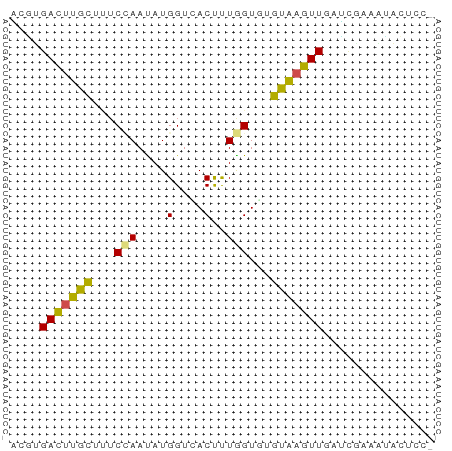
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:27:47 2011