
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,263,483 – 12,263,578 |
| Length | 95 |
| Max. P | 0.933126 |

| Location | 12,263,483 – 12,263,578 |
|---|---|
| Length | 95 |
| Sequences | 9 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 81.32 |
| Shannon entropy | 0.38852 |
| G+C content | 0.49977 |
| Mean single sequence MFE | -30.34 |
| Consensus MFE | -22.86 |
| Energy contribution | -22.04 |
| Covariance contribution | -0.81 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.38 |
| SVM RNA-class probability | 0.933126 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
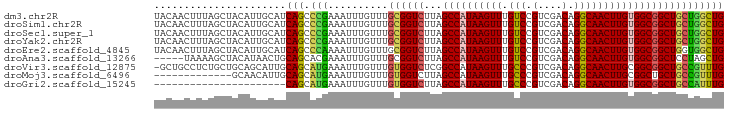
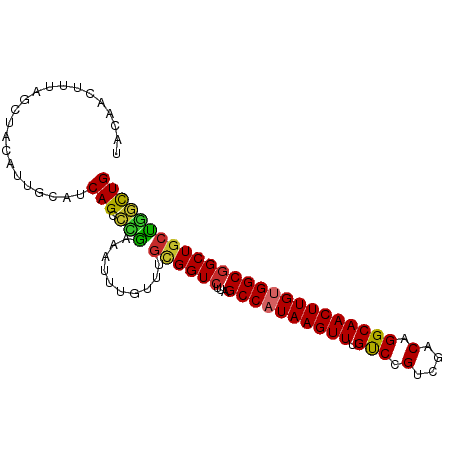
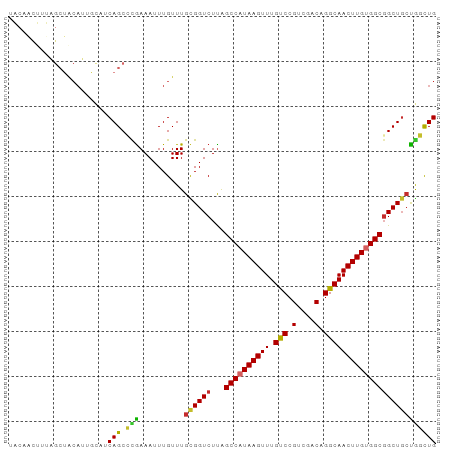
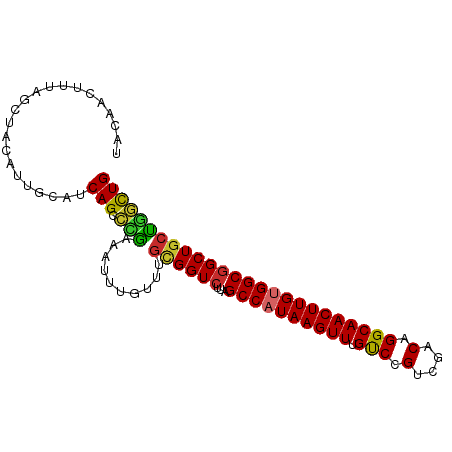
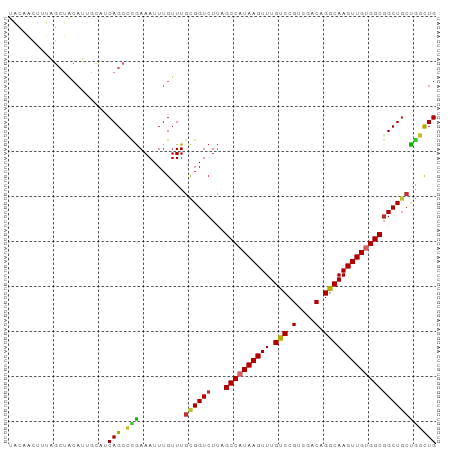
>dm3.chr2R 12263483 95 - 21146708 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droSim1.chr2R 10995159 95 - 19596830 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droSec1.super_1 9761156 95 - 14215200 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droYak2.chr2R 4417300 95 + 21139217 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droEre2.scaffold_4845 8992669 95 + 22589142 UACAACUUUAGCUACAUUGCAUCAGCCCAAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGGUGGCUG ........(((((((...((....(((((((.....)))).)))....((((((((((.(((.(....).))))))))))))).))..))))))) ( -32.70, z-score = -2.24, R) >droAna3.scaffold_13266 15639469 90 - 19884421 -----UAAAAGCUACAUAACUGCAGCACGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUCCUAGCUG -----....(((((....(((((((.((((...)))))))))))....((((((((((.(((.(....).)))))))))))))......))))). ( -30.20, z-score = -2.39, R) >droVir3.scaffold_12875 18767348 94 - 20611582 -GCUGCCUCUGCUGCAGCAUUGCAGCAUGAAAUUUGUUUGUGGUCUCGGCCAUAAGUUUGCCCGUCGACAGGCAACUUGCGGCGGCUGCCGUUUG -((((((((((((((......)))))).))...((((((((.(...(((.((......)).))).).)))))))).....))))))......... ( -36.20, z-score = -1.20, R) >droMoj3.scaffold_6496 12427689 82 - 26866924 -------------GCAACAUUGCAGCAUGAAAUUUGUUUGUGGUCUUAGCCAUAAGUUUGCCCGUCGACAGGCAACUUGCGGCUGCUGCCGUUUG -------------(((....)))((((.......))))...(((..(((((.((((((.(((.(....).))))))))).)))))..)))..... ( -24.20, z-score = -0.31, R) >droGri2.scaffold_15245 4846993 73 - 18325388 ----------------------CAGCAUGAAAUUUGUUUGUGGUCUUAGCCAUAAGUUUGCCCGUCGACAGGCAACUUGUGGCGGCUGCCAUUUG ----------------------.((((.......)))).(((((....((((((((((.(((.(....).)))))))))))))....)))))... ( -25.70, z-score = -2.31, R) >consensus UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ......................(((.(((..........((((((...((((((((((.(((.(....).))))))))))))))))))))))))) (-22.86 = -22.04 + -0.81)
| Location | 12,263,483 – 12,263,578 |
|---|---|
| Length | 95 |
| Sequences | 9 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 81.40 |
| Shannon entropy | 0.38775 |
| G+C content | 0.49911 |
| Mean single sequence MFE | -30.34 |
| Consensus MFE | -22.86 |
| Energy contribution | -22.04 |
| Covariance contribution | -0.81 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.930286 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
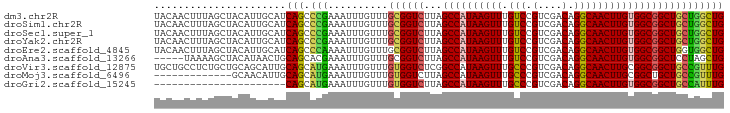
>dm3.chr2R 12263483 95 - 21146708 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droSim1.chr2R 10995159 95 - 19596830 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droSec1.super_1 9761156 95 - 14215200 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droYak2.chr2R 4417300 95 + 21139217 UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ..........((......))..(((((............((((((...((((((((((.(((.(....).))))))))))))))))))).))))) ( -31.02, z-score = -1.63, R) >droEre2.scaffold_4845 8992669 95 + 22589142 UACAACUUUAGCUACAUUGCAUCAGCCCAAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGGUGGCUG ........(((((((...((....(((((((.....)))).)))....((((((((((.(((.(....).))))))))))))).))..))))))) ( -32.70, z-score = -2.24, R) >droAna3.scaffold_13266 15639469 90 - 19884421 -----UAAAAGCUACAUAACUGCAGCACGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUCCUAGCUG -----....(((((....(((((((.((((...)))))))))))....((((((((((.(((.(....).)))))))))))))......))))). ( -30.20, z-score = -2.39, R) >droVir3.scaffold_12875 18767348 95 - 20611582 UGCUGCCUCUGCUGCAGCAUUGCAGCAUGAAAUUUGUUUGUGGUCUCGGCCAUAAGUUUGCCCGUCGACAGGCAACUUGCGGCGGCUGCCGUUUG .((((((((((((((......)))))).))...((((((((.(...(((.((......)).))).).)))))))).....))))))......... ( -36.20, z-score = -1.07, R) >droMoj3.scaffold_6496 12427689 82 - 26866924 -------------GCAACAUUGCAGCAUGAAAUUUGUUUGUGGUCUUAGCCAUAAGUUUGCCCGUCGACAGGCAACUUGCGGCUGCUGCCGUUUG -------------(((....)))((((.......))))...(((..(((((.((((((.(((.(....).))))))))).)))))..)))..... ( -24.20, z-score = -0.31, R) >droGri2.scaffold_15245 4846993 73 - 18325388 ----------------------CAGCAUGAAAUUUGUUUGUGGUCUUAGCCAUAAGUUUGCCCGUCGACAGGCAACUUGUGGCGGCUGCCAUUUG ----------------------.((((.......)))).(((((....((((((((((.(((.(....).)))))))))))))....)))))... ( -25.70, z-score = -2.31, R) >consensus UACAACUUUAGCUACAUUGCAUCAGCCCGAAAUUUGUUUGCGGUCUUAGCCAUAAGUUUGUCCGUCGACAGGCAACUUGUGGCGGCUGCUGGCUG ......................(((.(((..........((((((...((((((((((.(((.(....).))))))))))))))))))))))))) (-22.86 = -22.04 + -0.81)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:27:10 2011