
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 12,210,411 – 12,210,505 |
| Length | 94 |
| Max. P | 0.820989 |

| Location | 12,210,411 – 12,210,505 |
|---|---|
| Length | 94 |
| Sequences | 9 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 61.43 |
| Shannon entropy | 0.80626 |
| G+C content | 0.52542 |
| Mean single sequence MFE | -15.66 |
| Consensus MFE | -6.24 |
| Energy contribution | -6.01 |
| Covariance contribution | -0.23 |
| Combinations/Pair | 1.67 |
| Mean z-score | -0.91 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.820989 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 12210411 94 - 21146708 --UGCAGUCUCUUCUACUCCAUCCUCGUCCUUAUCCUUGUCAACACUUGAUCCAUCCUCUGCCUUCAAACGCUUCCUAAGGCGUCUGGCUGCACCC --(((((((.............................((((.....)))).................(((((......)))))..)))))))... ( -15.90, z-score = -2.04, R) >droWil1.scaffold_180700 6027262 79 + 6630534 -----------UGCAAUUGAUUCCGGUUCACCAUCCAAGCCAUCCUCCUGGACACCCUCGGCUUUAACUCUUUUCCUUAGACGUCUGUCA------ -----------.....((((..((((((.........)))).(((....))).......))..))))............(((....))).------ ( -10.00, z-score = 0.70, R) >droPer1.super_2 4916983 72 + 9036312 --------------------GACCGCCGCCUCUUCGUUAUCAAUGCAUGCUCCAUCCUCAGCUUUUACGCGCUUUCUCAAGCGCCGCGCUGC---- --------------------....((.((......(....)...((..(((........)))......((((((....)))))).)))).))---- ( -12.60, z-score = 0.68, R) >dp4.chr3 15846325 72 + 19779522 --------------------GACCGCCACCUCUUCGUUAUCAAUACAUGCUCCAUCCUCAGCUUUUACGCGCUUUCUCAAGCGCCGCGCUGC---- --------------------...(((......................(((........)))......((((((....)))))).)))....---- ( -12.00, z-score = -0.66, R) >droAna3.scaffold_13266 15588146 96 - 19884421 AACUCCAAGCUCUCUAGAGGCUCUUCAUACUUCUCUUGCUCUGUAAUUGAUCCUUCUUCGGCUUUUACGCGUUUCCUUAGCCGUCGGGUCGUUCCA .....(((((.....(((((.........)))))...))).)).....(((((.....(((((...............)))))..)))))...... ( -16.56, z-score = -0.06, R) >droEre2.scaffold_4845 8937446 94 + 22589142 --UGAAGUCUCUUCUUCUCCAUCCUCGUCCUUAUCUUUGUCUAUAGAGGAUCCCUCCUCUGCCUUUAAACGCUUCCUAAGGCGUCUGGCUGCGCCC --.((((......))))........(((................((((((....))))))(((.....(((((......)))))..))).)))... ( -20.20, z-score = -1.81, R) >droYak2.chr2R 4363580 94 + 21139217 --UGAAGUCUCUUCUACUCCAUCCUCGUCGUUAUCCUUGUCUAUAGAGGAUCCUUCCUCUGCCUUCAAACGCUUCCUAAGGCGUCUGGCUGCGCCC --.((((...))))...........(((.((((...(((....(((((((....)))))))....)))(((((......))))).)))).)))... ( -19.90, z-score = -1.27, R) >droSec1.super_1 9709738 94 - 14215200 --UGCAGUCUCUUCUAAUCCAUCCUCGUCCUUAUCCUUGUCUACACUGGAUCCAUCCUCUGCCUUCAAACGCUUCCUAAGGCGUCUGGCUGCGCCC --.((((((...........................(((....((..(((....)))..))....)))(((((......)))))..)))))).... ( -18.50, z-score = -2.13, R) >droSim1.chr2R 10941986 94 - 19596830 --UGCAGUCUCUUCUAAUCCAUCCUCGUCCUUAUCCUUGUCUACACUGCAUCCAUCCUCUGCCUUCAAACGCUUCCUAAGGCGUCUGGCUGCGCCC --.((((((......................................(((.........)))......(((((......)))))..)))))).... ( -15.30, z-score = -1.63, R) >consensus __UGCAGUCUCUUCUACUCCAUCCUCGUCCUUAUCCUUGUCUAUACAGGAUCCAUCCUCUGCCUUCAAACGCUUCCUAAGGCGUCUGGCUGCGCCC ............................................................(((.....((((((....))))))..)))....... ( -6.24 = -6.01 + -0.23)

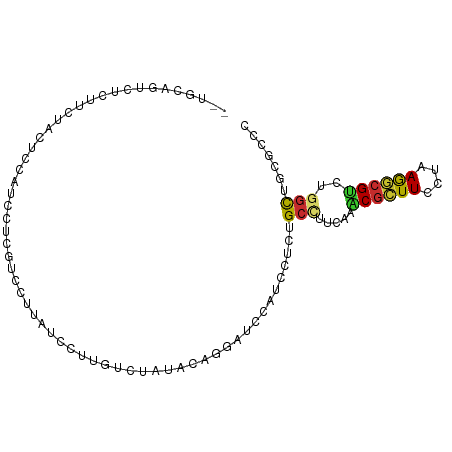
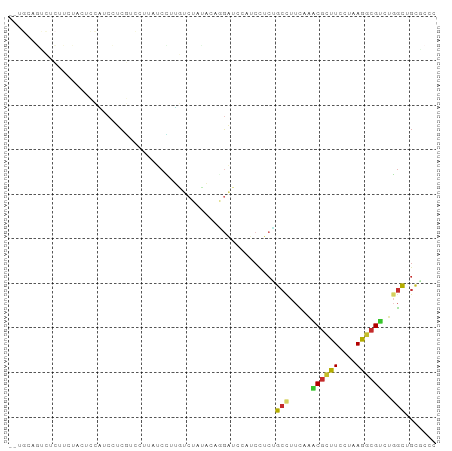
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:26:57 2011