
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,925,536 – 11,925,646 |
| Length | 110 |
| Max. P | 0.532274 |

| Location | 11,925,536 – 11,925,646 |
|---|---|
| Length | 110 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 71.70 |
| Shannon entropy | 0.55345 |
| G+C content | 0.60266 |
| Mean single sequence MFE | -38.50 |
| Consensus MFE | -20.90 |
| Energy contribution | -23.80 |
| Covariance contribution | 2.90 |
| Combinations/Pair | 1.32 |
| Mean z-score | -0.86 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.532274 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 11925536 110 + 21146708 GUGUUCUCUGGUACUUUCAGGUGCUGGGACUCGGGCAC---AAGGACUGGCGGCAGGACUAGGAGCAACAGUCGCGGUGGCAACGUCGAGACGCAUGCACGCUCCAUGCUCGC ..(((((..((((((....))))))))))).((((((.---.((..(((....)))..)).(((((....(((.((..(....)..)).)))(....)..))))).)))))). ( -42.30, z-score = -0.71, R) >droSim1.chr2R 10664162 113 + 19596830 GUGUGCUCUGGUACUUUCAGGUGCUGGAACUCGGGCACCACAAGGACCGGCGGCAGGACUAGGAGCAACAGUCGCGGUGGCAACGUCGAGACGCAUGCACGCUCCAUGCUCGC .(.(((((((((.((((..((((((.(....).))))))..))))))))).)))).)....(((((....(((.((..(....)..)).)))(....)..)))))........ ( -45.40, z-score = -0.87, R) >droSec1.super_1 9424525 110 + 14215200 GUGUGCUCUGGUACUUUCAGGUGCUGGAACUCGGGCAC---AAGGACUGGCGGCAGGACUAGGAGCAACAGUCGCGGUGGCAACGUCGAGACGCAUGCACGCUCCAUGCUCGC ......((..(((((....)))))..))...((((((.---.((..(((....)))..)).(((((....(((.((..(....)..)).)))(....)..))))).)))))). ( -42.70, z-score = -0.57, R) >droYak2.chr2R 4082144 104 - 21139217 ------GCUGGUACUUUCAGGUGCUGGGACUCGGGCAC---UGGGACUGGCGGCAGGACUAGGAGCAACAGUCGCGGAGGCAACGUCGAGACGCAUGCACGCUCCAUGCUCGC ------.(..(((((....)))))..)....((((((.---(((..(((....)))..)))(((((....(((.((..(....)..)).)))(....)..))))).)))))). ( -41.20, z-score = -0.90, R) >droEre2.scaffold_4845 8656548 104 - 22589142 ------GCUGGUACUUUCAGGUGCUGGGACUCGGGCAC---AAGGACUGGCGGCAGGACUAGGAGCAACAGUCGCGGAGGCAACGUCGAGACGCAUGCACGCUCCAUGCUCGC ------.(..(((((....)))))..)....((((((.---.((..(((....)))..)).(((((....(((.((..(....)..)).)))(....)..))))).)))))). ( -40.30, z-score = -1.05, R) >droAna3.scaffold_13266 4414710 88 - 19884421 --------------------GUAUUUCAGGUGGCGCCC-----CAGUGAACAGUGGGAGCAGGAGCUACAGUAGCGAAGGCAACCUCGAGACGCAUGCAUGCUCCAUGCUCGC --------------------.........(((((((((-----((.(....).)))).)).(((((....((..(((.(....).)))..))(....)..)))))..)).))) ( -29.40, z-score = -0.26, R) >droGri2.scaffold_15245 16886234 95 + 18325388 ---------------GCUUGGUUUUUCCUCUUUUUUUUGG-GGGGUUUUCUUAUAGGAACAACAA--GCAGUCGCAGCGGCAACGCCGAGACGCAUGUACGCUCCAUGUUCAA ---------------(((((......(((((.......))-)))(((((......)))))..)))--)).(((.(.(((....))).).)))(((((.......))))).... ( -28.20, z-score = -1.65, R) >consensus ______UCUGGUACUUUCAGGUGCUGGAACUCGGGCAC___AAGGACUGGCGGCAGGACUAGGAGCAACAGUCGCGGAGGCAACGUCGAGACGCAUGCACGCUCCAUGCUCGC .......((((((((....))))))))....(((((......((..(((....)))..)).(((((....(((.(((.(....).))).)))(....)..)))))..))))). (-20.90 = -23.80 + 2.90)

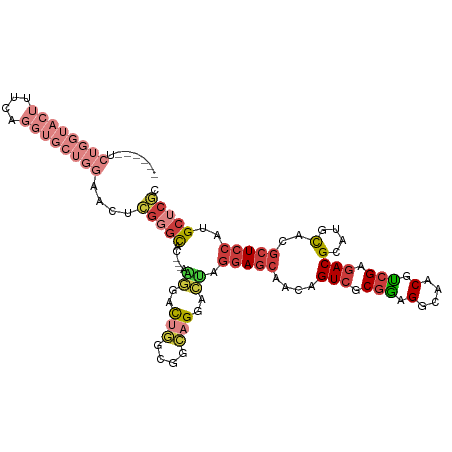

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:26:08 2011