
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 3,000,211 – 3,000,262 |
| Length | 51 |
| Max. P | 0.960542 |

| Location | 3,000,211 – 3,000,262 |
|---|---|
| Length | 51 |
| Sequences | 6 |
| Columns | 52 |
| Reading direction | forward |
| Mean pairwise identity | 88.96 |
| Shannon entropy | 0.20939 |
| G+C content | 0.54248 |
| Mean single sequence MFE | -16.35 |
| Consensus MFE | -11.95 |
| Energy contribution | -12.78 |
| Covariance contribution | 0.83 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.689395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
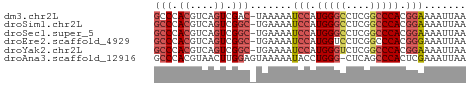
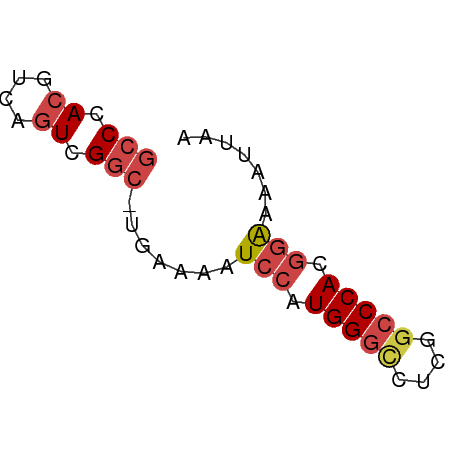
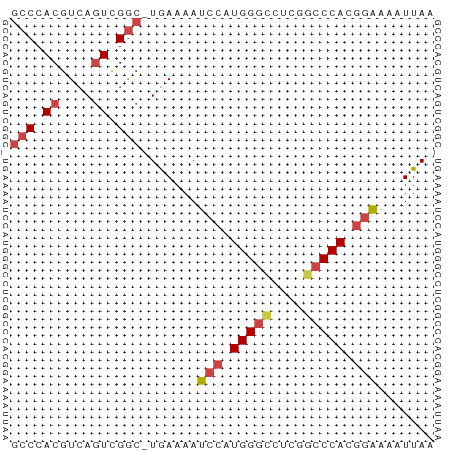
>dm3.chr2L 3000211 51 + 23011544 GCCCACGUCAGUCGAC-UAAAAAUCCAUGGGCCUCGGCCCACGGAAAAUUAA ......(((....)))-......(((.(((((....))))).)))....... ( -16.60, z-score = -2.82, R) >droSim1.chr2L 2959518 51 + 22036055 GCCCACGUCAGUCGGC-UGAAAAUCCAUGGGCCUCGGCCCACGGAAAAUUAA (((.((....)).)))-......(((.(((((....))))).)))....... ( -20.20, z-score = -2.88, R) >droSec1.super_5 1157319 51 + 5866729 GCCCACGUCAGUCGGC-UGAAAAUCCAUGGGCCUCGGCCCACGGAAAAUUAA (((.((....)).)))-......(((.(((((....))))).)))....... ( -20.20, z-score = -2.88, R) >droEre2.scaffold_4929 3046160 51 + 26641161 GCCCACGUCAGUCGGC-UGAAAAUCCAUGGUCCUCGGCCCACGGGAAAUUAA .(((((....)).(((-(((..((.....))..))))))...)))....... ( -13.00, z-score = -0.14, R) >droYak2.chr2L 2994969 51 + 22324452 GCCCACGUCAGUCGGC-UGAAAAUCCAUGGGUCUCGGCCCACGGAAAAUUAA (((.((....)).)))-......(((.(((((....))))).)))....... ( -18.10, z-score = -2.39, R) >droAna3.scaffold_12916 4422601 51 + 16180835 GCCCACGUAACUUGGAGUAAAAAUACCUGGG-CUCAGCCCACUCGAAAUUAA (((((.(((..((......))..))).))))-)................... ( -10.00, z-score = -0.84, R) >consensus GCCCACGUCAGUCGGC_UGAAAAUCCAUGGGCCUCGGCCCACGGAAAAUUAA (((.((....)).))).......(((.(((((....))))).)))....... (-11.95 = -12.78 + 0.83)
| Location | 3,000,211 – 3,000,262 |
|---|---|
| Length | 51 |
| Sequences | 6 |
| Columns | 52 |
| Reading direction | reverse |
| Mean pairwise identity | 88.96 |
| Shannon entropy | 0.20939 |
| G+C content | 0.54248 |
| Mean single sequence MFE | -19.35 |
| Consensus MFE | -15.47 |
| Energy contribution | -16.55 |
| Covariance contribution | 1.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.68 |
| SVM RNA-class probability | 0.960542 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
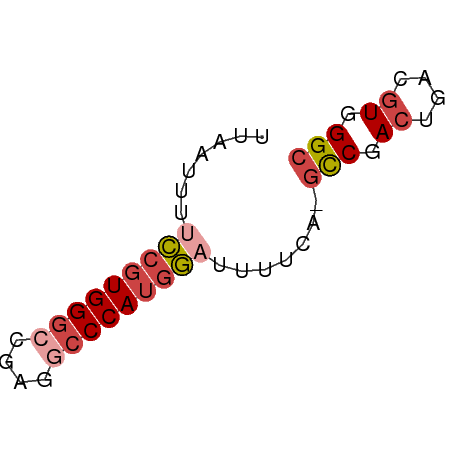
>dm3.chr2L 3000211 51 - 23011544 UUAAUUUUCCGUGGGCCGAGGCCCAUGGAUUUUUA-GUCGACUGACGUGGGC .(((...(((((((((....)))))))))...)))-(((.((....)).))) ( -21.80, z-score = -3.50, R) >droSim1.chr2L 2959518 51 - 22036055 UUAAUUUUCCGUGGGCCGAGGCCCAUGGAUUUUCA-GCCGACUGACGUGGGC .......(((((((((....)))))))))......-(((.((....)).))) ( -24.10, z-score = -3.75, R) >droSec1.super_5 1157319 51 - 5866729 UUAAUUUUCCGUGGGCCGAGGCCCAUGGAUUUUCA-GCCGACUGACGUGGGC .......(((((((((....)))))))))......-(((.((....)).))) ( -24.10, z-score = -3.75, R) >droEre2.scaffold_4929 3046160 51 - 26641161 UUAAUUUCCCGUGGGCCGAGGACCAUGGAUUUUCA-GCCGACUGACGUGGGC ........((((((.(....).)))))).......-(((.((....)).))) ( -15.70, z-score = -0.79, R) >droYak2.chr2L 2994969 51 - 22324452 UUAAUUUUCCGUGGGCCGAGACCCAUGGAUUUUCA-GCCGACUGACGUGGGC .......((((((((......))))))))......-(((.((....)).))) ( -19.10, z-score = -2.54, R) >droAna3.scaffold_12916 4422601 51 - 16180835 UUAAUUUCGAGUGGGCUGAG-CCCAGGUAUUUUUACUCCAAGUUACGUGGGC ...................(-((((.(((.(((......))).))).))))) ( -11.30, z-score = -0.23, R) >consensus UUAAUUUUCCGUGGGCCGAGGCCCAUGGAUUUUCA_GCCGACUGACGUGGGC .......(((((((((....))))))))).......(((.((....)).))) (-15.47 = -16.55 + 1.08)

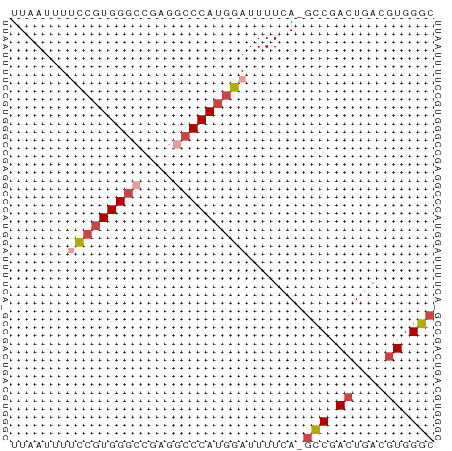
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:12:45 2011