
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,751,213 – 11,751,313 |
| Length | 100 |
| Max. P | 0.947842 |

| Location | 11,751,213 – 11,751,313 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 127 |
| Reading direction | forward |
| Mean pairwise identity | 82.17 |
| Shannon entropy | 0.29507 |
| G+C content | 0.54450 |
| Mean single sequence MFE | -38.75 |
| Consensus MFE | -29.90 |
| Energy contribution | -31.40 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.53 |
| SVM RNA-class probability | 0.947842 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
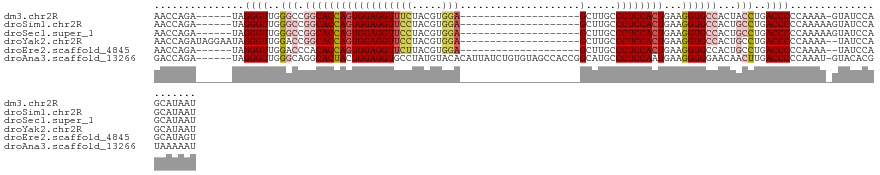
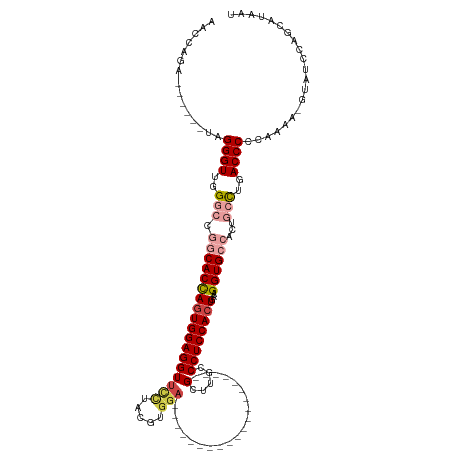
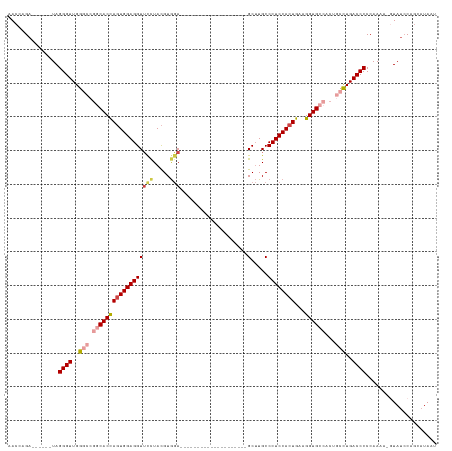
>dm3.chr2R 11751213 100 + 21146708 AACCAGA------UAGGGUUGGGCCGGCACCAGUGGAGGUUUCUACGUGGA--------------------GCUUGCCCUCCACUGAAGGUGCCACUACCUGACCCCCAAAA-GUAUCCAGCAUAAU .......------..((((..((..((((((.((((......))))(((((--------------------(......))))))....))))))....))..))))......-.............. ( -38.00, z-score = -2.06, R) >droSim1.chr2R 10493115 101 + 19596830 AACCAGA------UAGGGUUGGGCCGGCACCAGUGGAGGUUCCUACGUGGA--------------------GCUUGCCCUCCACUGAAGGUGCCACUGCCUGACCCCCAAAAAGUAUCCAGCAUAAU .......------..((((..(((.((((((((((((((((((.....)))--------------------).....))))))))...))))))...)))..))))..................... ( -43.60, z-score = -3.32, R) >droSec1.super_1 9245727 101 + 14215200 AACCAGA------UAGGGUUGGGCCGGCACCAGUGGAGGUUCCUACGUGGA--------------------GCUUGCCCUCCACUGAAGGUGCCACUGCCUGACCCCCAAAAAGUAUCCAGCAUAAU .......------..((((..(((.((((((((((((((((((.....)))--------------------).....))))))))...))))))...)))..))))..................... ( -43.60, z-score = -3.32, R) >droYak2.chr2R 3907324 105 - 21139217 AACCAGAUAGGAAUAGGGUUGGACCGGCACCAGUGGAGGUUCCUACGUGGA--------------------GCUUGCCCUCCACUGAAGGUGCCACUGCCUGACCCCCAAAA--UAUCCAGCAUAAU .....((((......(((((((...((((((((((((((((((.....)))--------------------).....))))))))...))))))....)).)))))......--))))......... ( -39.20, z-score = -2.34, R) >droEre2.scaffold_4845 8481943 99 - 22589142 AACCAGA------UAGGGUUGGACCCACACCAGUGGAGGUUCUUACGUGGA--------------------GCUUGCCCUCCACUGAAGGUGCCACUGCCUGACCCCCAAAA--UAUCCAGCAUAGU ..((...------..))((((((.((((....)))).((((.....(((((--------------------(......))))))...((((......)))))))).......--..))))))..... ( -29.80, z-score = -0.55, R) >droAna3.scaffold_13266 7440600 120 - 19884421 GACCAGA------UAGGGUUGGGCAGGCACUACUGGAGGUGCCUAUGUACACAUUAUCUGUGUAGCCACCGGCAUGCCCUCCAAUGAAGGUGGAACAACUUGACCCCCAAAU-GUACACGUAAAAAU .......------..((((..((.(((((((......)))))))...((((((.....)))))).(((((..(((........)))..))))).....))..))))......-.............. ( -38.30, z-score = -1.00, R) >consensus AACCAGA______UAGGGUUGGGCCGGCACCAGUGGAGGUUCCUACGUGGA____________________GCUUGCCCUCCACUGAAGGUGCCACUGCCUGACCCCCAAAA_GUAUCCAGCAUAAU ...............((((..(((.((((((((((((((.(((.....)))....................(....)))))))))...))))))...)))..))))..................... (-29.90 = -31.40 + 1.50)
| Location | 11,751,213 – 11,751,313 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 127 |
| Reading direction | reverse |
| Mean pairwise identity | 82.17 |
| Shannon entropy | 0.29507 |
| G+C content | 0.54450 |
| Mean single sequence MFE | -41.53 |
| Consensus MFE | -35.04 |
| Energy contribution | -35.27 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.11 |
| SVM RNA-class probability | 0.893983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
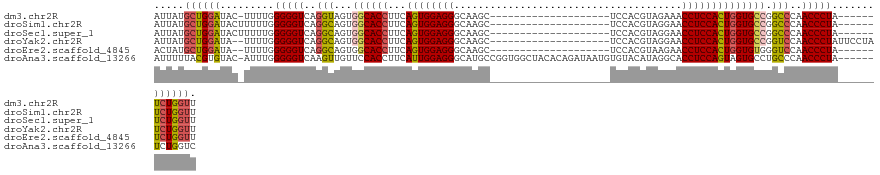
>dm3.chr2R 11751213 100 - 21146708 AUUAUGCUGGAUAC-UUUUGGGGGUCAGGUAGUGGCACCUUCAGUGGAGGGCAAGC--------------------UCCACGUAGAAACCUCCACUGGUGCCGGCCCAACCCUA------UCUGGUU .....((..(((.(-....)(((((..(((...((((((...((((((((....((--------------------.....)).....)))))))))))))).)))..))))))------))..)). ( -41.40, z-score = -1.87, R) >droSim1.chr2R 10493115 101 - 19596830 AUUAUGCUGGAUACUUUUUGGGGGUCAGGCAGUGGCACCUUCAGUGGAGGGCAAGC--------------------UCCACGUAGGAACCUCCACUGGUGCCGGCCCAACCCUA------UCUGGUU .....((..((((((....))((((..(((...((((((...((((((((......--------------------(((.....))).)))))))))))))).)))..))))))------))..)). ( -44.90, z-score = -2.38, R) >droSec1.super_1 9245727 101 - 14215200 AUUAUGCUGGAUACUUUUUGGGGGUCAGGCAGUGGCACCUUCAGUGGAGGGCAAGC--------------------UCCACGUAGGAACCUCCACUGGUGCCGGCCCAACCCUA------UCUGGUU .....((..((((((....))((((..(((...((((((...((((((((......--------------------(((.....))).)))))))))))))).)))..))))))------))..)). ( -44.90, z-score = -2.38, R) >droYak2.chr2R 3907324 105 + 21139217 AUUAUGCUGGAUA--UUUUGGGGGUCAGGCAGUGGCACCUUCAGUGGAGGGCAAGC--------------------UCCACGUAGGAACCUCCACUGGUGCCGGUCCAACCCUAUUCCUAUCUGGUU .....((..((((--.....(((((..(((...((((((...((((((((......--------------------(((.....))).)))))))))))))).)))..))))).....))))..)). ( -45.70, z-score = -2.67, R) >droEre2.scaffold_4845 8481943 99 + 22589142 ACUAUGCUGGAUA--UUUUGGGGGUCAGGCAGUGGCACCUUCAGUGGAGGGCAAGC--------------------UCCACGUAAGAACCUCCACUGGUGUGGGUCCAACCCUA------UCUGGUU .....((..(((.--.....(((((..(((....(((((...((((((((....((--------------------.....)).....)))))))))))))..)))..))))))------))..)). ( -35.10, z-score = -0.15, R) >droAna3.scaffold_13266 7440600 120 + 19884421 AUUUUUACGUGUAC-AUUUGGGGGUCAAGUUGUUCCACCUUCAUUGGAGGGCAUGCCGGUGGCUACACAGAUAAUGUGUACAUAGGCACCUCCAGUAGUGCCUGCCCAACCCUA------UCUGGUC .............(-(..(((((..........((((.......))))((((..(((...)))((((((.....))))))...((((((........))))))))))..)))))------..))... ( -37.20, z-score = -0.15, R) >consensus AUUAUGCUGGAUAC_UUUUGGGGGUCAGGCAGUGGCACCUUCAGUGGAGGGCAAGC____________________UCCACGUAGGAACCUCCACUGGUGCCGGCCCAACCCUA______UCUGGUU .....((((((.........(((((..(((...((((((...((((((((......................................)))))))))))))).)))..))))).......)))))). (-35.04 = -35.27 + 0.23)
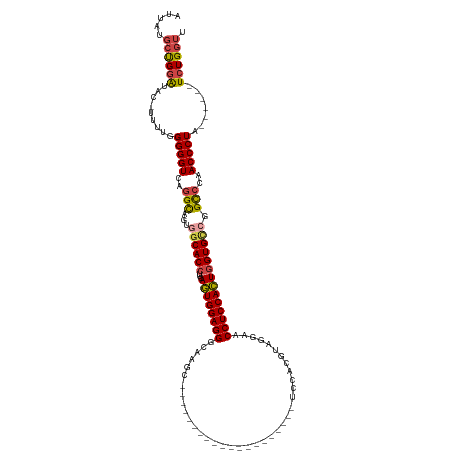
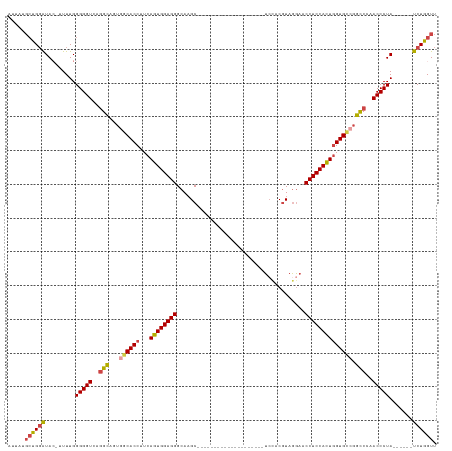
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:25:49 2011