
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,738,178 – 11,738,255 |
| Length | 77 |
| Max. P | 0.669576 |

| Location | 11,738,178 – 11,738,255 |
|---|---|
| Length | 77 |
| Sequences | 4 |
| Columns | 80 |
| Reading direction | forward |
| Mean pairwise identity | 74.84 |
| Shannon entropy | 0.41201 |
| G+C content | 0.62370 |
| Mean single sequence MFE | -19.86 |
| Consensus MFE | -10.16 |
| Energy contribution | -11.29 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.587923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
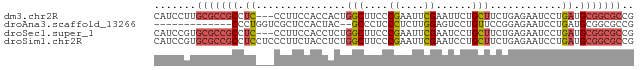
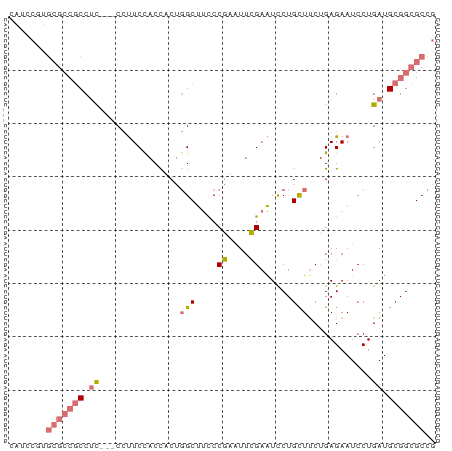
>dm3.chr2R 11738178 77 + 21146708 CAUCCUUGCGCCGCCUC---CCUUCCACCACUGGCUUCCCGAAUUCGAAUUCUGCUUCUGAGAAUCCUGAUGCGGCGCCG .......(((((((.((---........((..(((....((....))......)))..))........)).))))))).. ( -19.49, z-score = -2.05, R) >droAna3.scaffold_13266 7426891 65 - 19884421 -------------CCCUGGUCGCUCCACUAC--GCCCUCCCUCUUGGAGUCCUGUUCCGGAGAAUCCUGAUGCGGCGCCG -------------....((.((((.((....--(..((((.....))))..)...((.((.....)).)))).)))))). ( -15.80, z-score = 0.44, R) >droSec1.super_1 9232401 77 + 14215200 CAUCCGUGCGCCGCCUC---CCUUCCACCUCUGGCUUCCCGAAUUCGAAUCCUGCUUCUGAGAAUCCUGAUGCGGCGCCG ....((.(((((((.((---........(((.(((....((....))......)))...)))......)).))))))))) ( -22.04, z-score = -2.56, R) >droSim1.chr2R 10477680 80 + 19596830 CAUCCGUGCGCCGCCUCCUCCCUUCUACCUCUGGCUUCCCGAAUUCGAAUCCUGCUUCUGAGAAUCCUGAUGCGGCGCCG ....((.(((((((.((.....((((......(((....((....))......)))....))))....)).))))))))) ( -22.10, z-score = -2.86, R) >consensus CAUCCGUGCGCCGCCUC___CCUUCCACCACUGGCUUCCCGAAUUCGAAUCCUGCUUCUGAGAAUCCUGAUGCGGCGCCG .......(((((((.((.....(((.......((.(((........))).)).......)))......)).))))))).. (-10.16 = -11.29 + 1.13)

| Location | 11,738,178 – 11,738,255 |
|---|---|
| Length | 77 |
| Sequences | 4 |
| Columns | 80 |
| Reading direction | reverse |
| Mean pairwise identity | 74.84 |
| Shannon entropy | 0.41201 |
| G+C content | 0.62370 |
| Mean single sequence MFE | -25.30 |
| Consensus MFE | -15.20 |
| Energy contribution | -17.07 |
| Covariance contribution | 1.88 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.669576 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
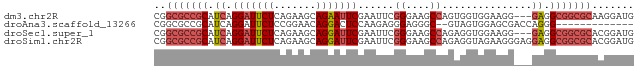
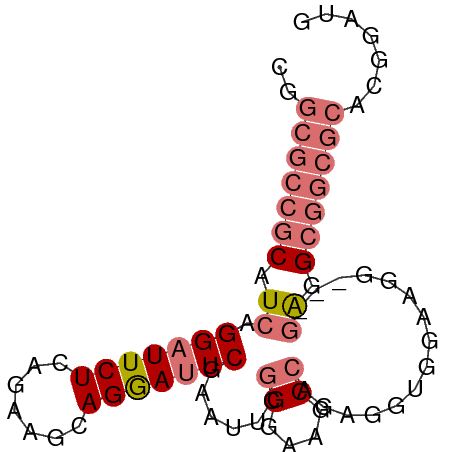
>dm3.chr2R 11738178 77 - 21146708 CGGCGCCGCAUCAGGAUUCUCAGAAGCAGAAUUCGAAUUCGGGAAGCCAGUGGUGGAAGG---GAGGCGGCGCAAGGAUG ..(((((((.((.((((((((....).)))))))......((....))............---)).)))))))....... ( -26.70, z-score = -1.99, R) >droAna3.scaffold_13266 7426891 65 + 19884421 CGGCGCCGCAUCAGGAUUCUCCGGAACAGGACUCCAAGAGGGAGGGC--GUAGUGGAGCGACCAGGG------------- ..(((((............(((......)))((((.....)))))))--))..(((.....)))...------------- ( -18.70, z-score = 0.61, R) >droSec1.super_1 9232401 77 - 14215200 CGGCGCCGCAUCAGGAUUCUCAGAAGCAGGAUUCGAAUUCGGGAAGCCAGAGGUGGAAGG---GAGGCGGCGCACGGAUG (((((((((.((.((((((((....).)))))))...(((((....)).)))........---)).))))))).)).... ( -27.90, z-score = -2.21, R) >droSim1.chr2R 10477680 80 - 19596830 CGGCGCCGCAUCAGGAUUCUCAGAAGCAGGAUUCGAAUUCGGGAAGCCAGAGGUAGAAGGGAGGAGGCGGCGCACGGAUG (((((((((.((.((((((((....).)))))))...(((((....)).)))...........)).))))))).)).... ( -27.90, z-score = -2.43, R) >consensus CGGCGCCGCAUCAGGAUUCUCAGAAGCAGGAUUCGAAUUCGGGAAGCCAGAGGUGGAAGG___GAGGCGGCGCACGGAUG ..(((((((.((.(((((((.......)))))))......((....))...............)).)))))))....... (-15.20 = -17.07 + 1.88)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:25:46 2011