
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,715,426 – 11,715,546 |
| Length | 120 |
| Max. P | 0.500000 |

| Location | 11,715,426 – 11,715,546 |
|---|---|
| Length | 120 |
| Sequences | 14 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.31 |
| Shannon entropy | 0.40258 |
| G+C content | 0.57143 |
| Mean single sequence MFE | -46.10 |
| Consensus MFE | -21.38 |
| Energy contribution | -21.43 |
| Covariance contribution | 0.05 |
| Combinations/Pair | 1.58 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
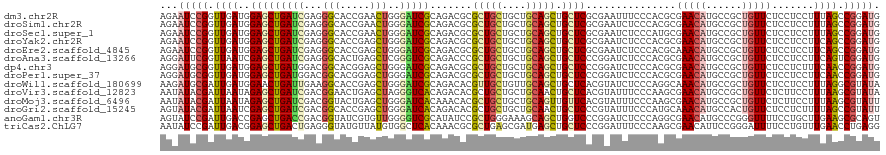
>dm3.chr2R 11715426 120 + 21146708 AGAAUCCGGUUGAUGGAGCUGAUCGAGGGCACCGAACUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCGCGAAUUUCCCACGCGAACAUGCCGCUGUUCUCCUCCUUUAGCCGGAUG ...((((((((((.((((..((.(.((((((.((...(((((((((.((.(((((((....))))).))))))))...)))))..))....)))).))).))..)))).)))))))))). ( -55.20, z-score = -3.84, R) >droSim1.chr2R 10455189 120 + 19596830 AGAAUCCGGUUGAUGGAGCUGAUCGAGGGCACCGAACUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCGCGAAUCUCCCACGCGAACAUGCCGCUGUUCUCCUCCUUUAGCCGGAUG ...((((((((((.((((..((.(.((((((.((...(((((((((.((.(((((((....))))).))))))))...)))))..))....)))).))).))..)))).)))))))))). ( -55.20, z-score = -3.65, R) >droSec1.super_1 9209615 120 + 14215200 AGAAUCCGGUUGAUGGAGCUGAUCGAGGGCACCGAACUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCGCGAAUCUCCCAUGCGAACAUGCCGCUGUUCUCCUCCUUUAGCCGGAUG ...((((((((((.((((..((.(.((((((.((...(((((((((.((.(((((((....))))).))))))))...)))))..))....)))).))).))..)))).)))))))))). ( -55.20, z-score = -3.73, R) >droYak2.chr2R 3869216 120 - 21139217 AGAAUCCGGUUGAUGGAGCUGAUCGAGGGCACCGAGCUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCGCGAAUCUCCCACGCGAACAUGCCGCUGUUCUCCUCCUUCAGCCGGAUG ...((((((((((.((((..((.(.((((((....(((((((((((.((.(((((((....))))).))))))))...))))).)).....)))).))).))..)))).)))))))))). ( -60.10, z-score = -3.97, R) >droEre2.scaffold_4845 8444947 120 - 22589142 AGAAUCCGGUUGAUGGAGCUGAUCGAGGGCACCGAGCUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCGCGAAUCUCCCACGCAAACAUGCCGCUGUUCUCCUCCUUCAGCCGGAUG ...((((((((((.((((..((.(.((((((....(((((((((((.((.(((((((....))))).))))))))...))))).)).....)))).))).))..)))).)))))))))). ( -59.00, z-score = -4.01, R) >droAna3.scaffold_13266 7405669 120 - 19884421 AGGAUUCGGUUAAUCGAGCUGAUCGAGGGCACUGAGCUCGGGUCGCAGACCCGCUGCUGCUGCAGCUGCUCCCGGAUCUCCCACGCGAACAUGCCGCUGUUCUCCUCCUUCAGUCGGAUG .............((((.((((..((((((...((((..(((((...)))))(((((....))))).))))..((.....))..))(((((......))))).))))..))))))))... ( -44.80, z-score = -0.29, R) >dp4.chr3 309080 120 - 19779522 AGGAUGCGGUUGAUGGAGCUGAUGGACGGCACGGAGCUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCCCGGAUCUCCCACGCGAACAUGCCGCUGUUCUCCUCUUUCAACCGGAUG ...((.(((((((.((((..((.(((((((..((((((((((.(((....))(((((....))))).).))))))..))))...((......)).))))))))))))).))))))).)). ( -53.10, z-score = -2.28, R) >droPer1.super_37 288104 120 - 760358 AGGAUGCGGUUGAUGGAGCUGAUGGACGGCACGGAGCUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCCCGGAUCUCCCACGCGAACAUGCCGCUGUUCUCCUCCUUCAACCGGAUG ...((.(((((((.((((..((.(((((((..((((((((((.(((....))(((((....))))).).))))))..))))...((......)).))))))))))))).))))))).)). ( -54.90, z-score = -2.62, R) >droWil1.scaffold_180699 1739525 120 - 2593675 AAGAUGCGAUUGAUGGAACUGAUUGAAGGCACCGAGCUGGGAUCGCAGACACGUUGCUGUUGCAGCUGCUCACGUAUCUCCCAGGCAAACAUGCCGCUGUUCUCCUCCUUUAGGCGUAUA ...(((((.((((.(((...((.....((((....(((((((.((.((.((.(((((....)))))))))..))....))))).)).....)))).....))...))).)))).))))). ( -34.20, z-score = 0.65, R) >droVir3.scaffold_12823 2270059 120 - 2474545 AAUAUACGAUUAAUAGAGCUGAUCGACGGAACUGAGCUAGGGUCACAGACACGCUGCUGCUGCAACUGCUCACGUAUUUCCCAAGCGAACAUGCCGCUGUUCUCUUCCUUUAAGCGUAUA ..((((((.((((..(((..((.((.(((......(((.(((......((..((....)).((....))....))....))).))).......))).)).))..)))..)))).)))))) ( -28.02, z-score = 0.32, R) >droMoj3.scaffold_6496 26814669 120 - 26866924 AAUAUACGAUUAAUAGAGCUGAUCGACGGUACUGAGCUGGGAUCACAAACACGCUGCUGCUGCAGUUGUUCACGUAUUUCCCAAGCGAACAUGCCGCUGUUCUCUUCCUUUAAGCGUAUU ...(((((.((((..(((..((.((.(((((.((.(((((((..((.((((.(((((....)))))))))...))...)))).)))...))))))).)).))..)))..)))).))))). ( -35.90, z-score = -2.20, R) >droGri2.scaffold_15245 6450658 120 + 18325388 AGUAUACGAUUAAUCGAGCUGAUCGACGGCACCGAGCUGGGAUCACAGACACGCUGCUGCUGCAACUGCUCCCGUAUUUCCCAUGCAAACAUGCCACUGUUCUCCUCUUUUAGCCGUAUU ...(((((..(((..(((..((.((..((((....(((((((..((.((((.(.(((....))).))).))..))...))))).)).....))))..)).))..)))..)))..))))). ( -29.90, z-score = -0.55, R) >anoGam1.chr3R 34546101 120 - 53272125 AGUAUCCGAUUGACCGAGCUGACCGACGGUAUCGUGUUGGGGUCGCAUAUCCGCUGGGAAAGCAGCUGGUCCCGGAUCUCCCAGGCGAACAUGCCCGGGUUUUCCUGCUUGAAGCGCAGU ((.(((((...((((.(((((.(((.((((((.(((.......)))))).))).))).....))))))))).)))))))(((.(((......))).))).....((((.......)))). ( -42.10, z-score = 0.25, R) >triCas2.ChLG7 15303404 120 + 17478683 AAUAUCCGAUUGACGGAGCUGACUGAGGGUAUGUUAUGUGGCUCACAAACGCGCUGAGCGAUGAGCUGCUCCCGGAUUUCCCAAGCGAACAUUCCGGGAUUUUCCUGUUUGAACCUGAGG ....((((.....)))).....((.(((.........(..(((((....(((.....))).)))))..)(((((((.(((......)))...)))))))..............))).)). ( -37.80, z-score = -1.29, R) >consensus AGAAUCCGGUUGAUGGAGCUGAUCGAGGGCACCGAGCUGGGAUCGCAGACGCGCUGCUGCUGCAGCUGCUCCCGAAUCUCCCACGCGAACAUGCCGCUGUUCUCCUCCUUUAGCCGGAUG ...(((((.((((..(((((((((...(((.....)))..))))).......(((((....))))).))))...............(((((......))))).......)))).))))). (-21.38 = -21.43 + 0.05)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:25:36 2011