
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,590,255 – 11,590,346 |
| Length | 91 |
| Max. P | 0.709980 |

| Location | 11,590,255 – 11,590,346 |
|---|---|
| Length | 91 |
| Sequences | 11 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 76.72 |
| Shannon entropy | 0.50584 |
| G+C content | 0.50139 |
| Mean single sequence MFE | -22.60 |
| Consensus MFE | -13.90 |
| Energy contribution | -13.82 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.03 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.709980 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 11590255 91 + 21146708 UUGUCGUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCACCAUCUGGCGACGAUUUGUUGGCAGCAGCGCACCGCCGCCAGACGCC- ..(((((((((.((((.(((..(((((....)))))))).))))......)))))))))...(((((.((........)).))))).....- ( -29.70, z-score = -1.28, R) >droSim1.chr2R 10324638 91 + 19596830 UUGUCGUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGCCAUCUGGCGACGAUUUGUUGGCAGCAGCGCAUAGCCAUCCGACGCC- ....(((((...((((((...(((..((((....((((((((((.....)))))))))).)))).((....)))))))))))..)))))..- ( -29.90, z-score = -1.54, R) >droSec1.super_1 9083624 91 + 14215200 UUGCCGUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGCCAUCUGGCGACGAUUUGUUGGCAGCAGCGCACCGUCGUUAGACCAC- (((((((.((....)).)).......((((....((((((((((.....)))))))))).))))))))).........(((....)))...- ( -26.30, z-score = -0.40, R) >droYak2.chr2R 3740243 69 - 21139217 UUGUCGUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGCCAUCUGGCGACGAUUUGUUGGCAG----------------------- .((((((.((....)).)).......((((....((((((((((.....)))))))))).)))))))).----------------------- ( -21.20, z-score = -2.01, R) >droEre2.scaffold_4845 8313712 79 - 22589142 AUGUCGUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGCCAUCUGGCGACGAUUUGCUGGCAAC--CCAACCGCC----------- ..(((((((((.((((..(((.(((((....))))).)).)..))))...))))))))).....(((...--......)))----------- ( -22.70, z-score = -1.35, R) >droAna3.scaffold_13266 7279320 91 - 19884421 UUGUCCUUGCCUCGGUUACAUUGUAAAACUUUUUAUUGUUGCCGCCAGAUGGCGACGAUUUGUUGGCAGCUCUUCACCUUCCUUAG-CCACC .............(((((....(((((....))))).(((((((.((((((....).))))).))))))).............)))-))... ( -19.40, z-score = -0.69, R) >dp4.chr3 179753 88 - 19779522 UUCUUCUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGGCAUCUGGCGACGAUUUGUUGGUAAC---GCACCUCCAUCAACCCCC- .............((((.........((((....(((((((((((...))))))))))).))))(((...---..)))......))))...- ( -21.50, z-score = -1.51, R) >droPer1.super_37 159051 88 - 760358 UUCUUCUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGGCAUCUGGCGACGAUUUGUUGGUAAC---GCACCUCUAUCAACCCCC- .............((((.........((((....(((((((((((...))))))))))).))))(((...---..)))......))))...- ( -21.50, z-score = -1.72, R) >droVir3.scaffold_12823 936267 89 - 2474545 AUGUCGUCGCCUUAGUUACAUUGUAAAACGUUUUAUUGUCGCUGGCAGCUGGCGACGAUUUGUUGCCGCC--CC-AACUCCUCCAGCUGCUA ..((((((((((((((.(((..(((((....)))))))).))))).....))))))))).....((.((.--..-..........)).)).. ( -21.12, z-score = -0.27, R) >droMoj3.scaffold_6496 382964 84 + 26866924 AUGUCGUCGCCUUAGUUACAUUGUAAAACGUUUUAUUGUCGCUGGCAGCUGGCAACGAUUUGUUGCUGCC--AU-CCCUCCCCCAUC----- .............(((.(((..(((((....)))))))).)))((((((.(((((....)))))))))))--..-............----- ( -17.20, z-score = -0.59, R) >droGri2.scaffold_15245 5748238 81 + 18325388 AUGUCGUCGUCUUAGUUACAUUGUAAAACGUUUUAUUGUCGCUGGCAGCUGGCGACGAUUUGUUGCUGCC--CCCAACUGCUG--------- ............(((((.....(((.((((....((((((((..(...)..)))))))).))))..))).--...)))))...--------- ( -18.10, z-score = -0.02, R) >consensus UUGUCGUCGCCUUGGUUACAUUGUAAAACGUUUUAUUGUCGCCGCCAUCUGGCGACGAUUUGUUGGCAGC__CGCACCUCCAUCAG_C_CC_ ..........................((((....((((((((((.....)))))))))).))))............................ (-13.90 = -13.82 + -0.08)


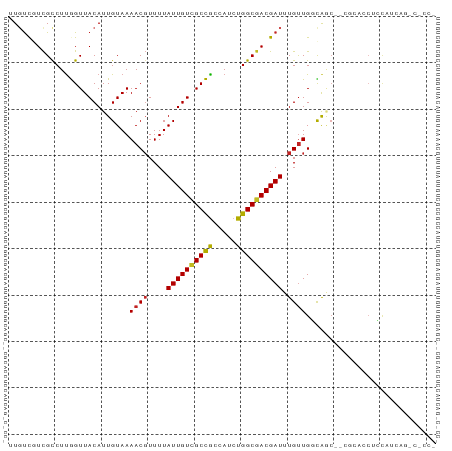
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:25:14 2011