
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,306,478 – 11,306,577 |
| Length | 99 |
| Max. P | 0.891035 |

| Location | 11,306,478 – 11,306,577 |
|---|---|
| Length | 99 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 66.18 |
| Shannon entropy | 0.66331 |
| G+C content | 0.48001 |
| Mean single sequence MFE | -20.69 |
| Consensus MFE | -7.66 |
| Energy contribution | -7.54 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.891035 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 11306478 99 - 21146708 CCUCCAUUCCCCCAAACCCCUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUGA------AGGCAGCU--CCCCAUUUGCCCCCCACG------- .....................(((..(((.....))).(((((((((.....((((......)))).....))------))))))).--........))).......------- ( -19.90, z-score = -2.64, R) >droAna3.scaffold_13266 7723203 89 - 19884421 --------UUCCUACCCUCUUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGCGA------AGGCAGCAC-UUCCACUCUCCCGCU---------- --------.............(((..(((.....))).(((((((((.((..((((......))))...))))------)))))))..-............)))---------- ( -18.00, z-score = -1.33, R) >droEre2.scaffold_4845 8019878 105 + 22589142 GCUCCAUUCCCGCCACCCCCUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUGGAGGCGGAGGCAGCU--CCCCAUUUGCCCCCCACG------- ..(((((((..((((.....)))).......(((((.........))))).................)))))))((.((.(((((..--......)))))))))...------- ( -26.20, z-score = -0.71, R) >droYak2.chr2R 3444814 84 + 21139217 -------UCUCCAUUCCCCUUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUGG------AGGCAGCU--UCCC-UCCACU-------------- -------...(((.......)))...............(((((((((.....((((......)))).....))------))))))).--....-......-------------- ( -17.60, z-score = -0.83, R) >droSec1.super_1 8786080 105 - 14215200 CCUCCAUUCCCG-CAACCCCUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUGG------AGGCAGCU--CCCCAUUUGCCCCCCCCCCCCCACG ...........(-(((...(((..........)))...(((((((((.....((((......)))).....))------))))))).--......))))............... ( -21.10, z-score = -1.81, R) >droSim1.chr2R 10033039 98 - 19596830 CCUCCAUUCCCG-CACCCCCUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUGG------AGGCAGCU--CCCCAUUUGCCCCCCACG------- ...........(-((....(((..........)))...(((((((((.....((((......)))).....))------))))))).--.......)))........------- ( -20.20, z-score = -1.19, R) >droWil1.scaffold_181141 2210203 94 + 5303230 -----AUCUGUUUUCUCCCCUCGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUUA------GAAUGGCA--GUGGAAGUGUGAGCAGAG------- -----.........(((..((((((..((.....))(((((((((((((.((((((.........))))))))------))).))))--))))..).)))))..)))------- ( -24.40, z-score = -1.82, R) >droGri2.scaffold_15245 7083395 92 + 18325388 CCUCCAUUCCUAAUGCCCCGCGAGCAAUCAAACAAAUCGCUGCCUCUCAUAUUUAUGCAUAAAUAAUGAG---------ACGAGGUGUGUUUUGCUUGGCU------------- ..............(((....((((((....(((..(((.....(((((((((((....))))).)))))---------))))..)))...))))))))).------------- ( -22.10, z-score = -1.73, R) >dp4.chr3 11945385 79 - 19779522 ----------------------GCGAAUCAAACAGAUCGCUGCCUCUUGAAUUUAUGCAUAAAUAAUGAACGC------AAGCGGCACUUUUCAUCCGUCCUCCAAG------- ----------------------((((.((.....))))))((((.((((..(((((.........)))))..)------))).))))....................------- ( -17.80, z-score = -2.10, R) >droPer1.super_44 157544 79 - 614636 ----------------------GCGAAUCAAACAGAUCGCUGCCUCUUGUAUUUAUGCAUAAAUAAUGAACGC------AAGCGGCACUUUUCAUCCGUCCUCCAAG------- ----------------------((((.((.....))))))((((.((((..(((((.........)))))..)------))).))))....................------- ( -19.60, z-score = -2.91, R) >consensus _____AUUCCCC_CACCCCCUGGCCAAUCAAACAGAUCGCUGCCUUUUGUAUUUAUGCAUAAAUAAUGAGUGG______AGGCAGCU__UCCCAUUUGCCCCCCACG_______ ......................................((((((((...((((((....))))))..)))...........)))))............................ ( -7.66 = -7.54 + -0.12)
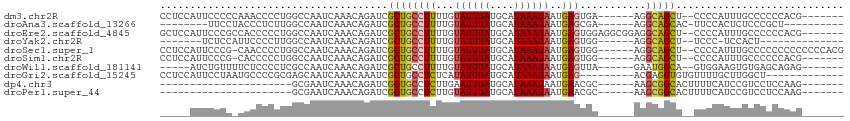
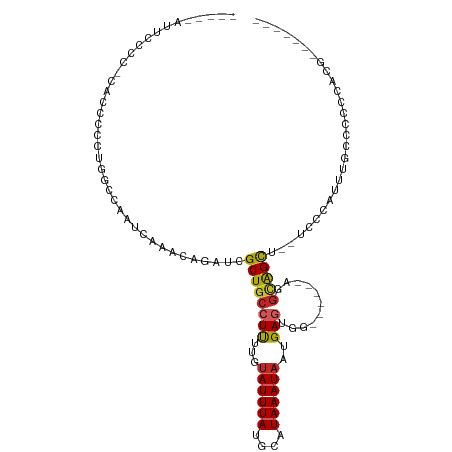

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:24:46 2011