
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 11,006,511 – 11,006,628 |
| Length | 117 |
| Max. P | 0.692576 |

| Location | 11,006,511 – 11,006,628 |
|---|---|
| Length | 117 |
| Sequences | 8 |
| Columns | 132 |
| Reading direction | reverse |
| Mean pairwise identity | 72.91 |
| Shannon entropy | 0.50373 |
| G+C content | 0.49416 |
| Mean single sequence MFE | -37.33 |
| Consensus MFE | -18.77 |
| Energy contribution | -17.65 |
| Covariance contribution | -1.12 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.692576 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 11006511 117 - 21146708 GGCGGCAUAUCCCGCCGAUUUUCGGCGUGGGCUGAUUUUUA--GCCGCAUUCGAUUUCGAUUUCGGGUUCAAGACUUGCCAUUCAACCUUUG--GUU----GGCUGGCCU-----AUAAA--UUUGCACUAU (((((((..((((((((.....))))).)((((((...)))--)))(.((((((........)))))).)..))..))))..((((((...)--)))----))...))).-----.....--.......... ( -37.60, z-score = -1.47, R) >droSim1.chr2R 9753871 116 - 19596830 ---GGCACAGCCCGCUGAUUUUCGGCGUGGGCUGAUUUUUA--GCCGCAUUCGAUUUCGAUUUCGGGUUCAAGACUUGCCAUUCAACCUUUGGGGUU----GGCUGGCCU-----AUAAA--UUUGCACUAU ---.(((((((((((((.....))))..))))))((((.((--((((...(((....)))...)))...........((((..(((((.....))))----)..))))))-----).)))--).)))..... ( -40.80, z-score = -2.45, R) >droSec1.super_1 8509349 116 - 14215200 ---GGCAUAGCCCGCCGAUUUUCGGCGUGGGCUGAUUUUUA--GCCGCAUUCGAUUUCGAUUUCGGGUUCAAGACUUGCCGUUCAACCUUUGGGGUU----GGCUGGCCU-----AUAAA--UUUGCACUAU ---.(((((((((((((.....))))..))))))((((.((--((((...(((....)))...)))...........((((..(((((.....))))----)..))))))-----).)))--).)))..... ( -40.30, z-score = -1.78, R) >droYak2.chr2R 10935087 108 - 21139217 ---GGCAUAGCCCGCCGAUUUUCGGCGUGGGCUGAUUUUUG--GCCGCA------UUCGAUUUCGGGUUCAAGACUUGCCAUUCAACCUUUG--AUU----GGCUGGCCU-----AUAAA--UUUGCGCUAU ---(((....(((((((.....))))).))((.(((((..(--((((.(------((((....))))).).......((((.(((.....))--).)----))).)))).-----..)))--)).))))).. ( -40.30, z-score = -2.50, R) >droEre2.scaffold_4845 7729640 108 + 22589142 ---GGCACAGCCCGCCGAUUUUCGGCGUGGGCUGAUUUUUA--GCCGCA------UUCGAUUUCGGGUUCAAGACUUGCCAUUCAACCUCUG--GUU----GGCUGGCCU-----AUAAA--UUGGCUCUAU ---....((((((((((.....))))..))))))......(--((((.(------((((....))))).).......((((..(((((...)--)))----)..))))..-----.....--..)))).... ( -39.80, z-score = -2.37, R) >droAna3.scaffold_13266 17551662 119 + 19884421 ---GGCCCAGCCCGCCAGUUUUUAGCGUGGGCUUUAUUUUA--GCCACA------UUCGAUUUGGGGUUCAAGACUUGCCAUUCAACCUCUGCCUCU--CCGGCGGGCCUGUUCCAUAAAUCUCUAUACUAU ---((..(((((((((........(.(((((((.......)--))).))------).).....((((....(((.(((.....)))..)))....))--)))))))).)))..))................. ( -35.20, z-score = -1.52, R) >dp4.chr3 8882896 125 - 19779522 ---GGCACUGGCCGCUGUUCAGUGGCGUGGGCUGAUGUCUGAUGUCCCACCCGAAUUUGAUUUAGGGUUCAAGUCUUGCCAUUCAACCUUUACCAUUGCGUACUCCCGUUGUGCUAUAAAAAACUAGU---- ---.(((.(((.....(((.((((((((((((...........).)))))..((((((......)))))).......)))))).))).....))).)))((((.......))))..............---- ( -31.10, z-score = 0.47, R) >droPer1.super_4 2768283 125 - 7162766 ---GGCACAGGCCGCUAUUUAGUGGCGUGGGCUGAUGUCUGAUGUGCCACCCGAAUUUGAUUUAGGGUUCAAGUCUUGCCAUUCAACCUUUACUAUUGCGUACUCCGGUUGUGCUAUAAAAAACUAGU---- ---((((((.((((((....))))))..((((....))))..))))))((((............)))).........((....(((((..(((......)))....))))).))..............---- ( -33.50, z-score = 0.01, R) >consensus ___GGCACAGCCCGCCGAUUUUCGGCGUGGGCUGAUUUUUA__GCCGCA__CGA_UUCGAUUUCGGGUUCAAGACUUGCCAUUCAACCUUUG__GUU____GGCUGGCCU_____AUAAA__UUUGCACUAU ...(((((((((((((((...)))))..))))))........................((........))......)))).....................((....))....................... (-18.77 = -17.65 + -1.12)

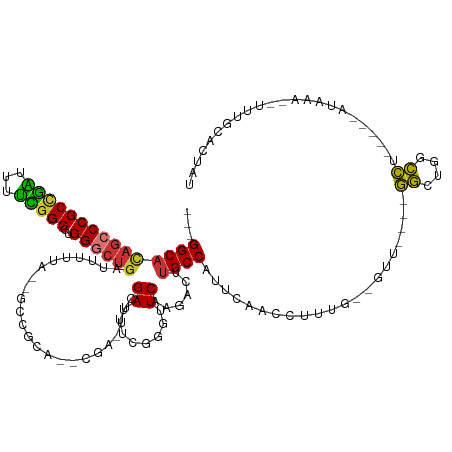

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:24:14 2011