
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 10,546,077 – 10,546,148 |
| Length | 71 |
| Max. P | 0.996978 |

| Location | 10,546,077 – 10,546,148 |
|---|---|
| Length | 71 |
| Sequences | 7 |
| Columns | 71 |
| Reading direction | forward |
| Mean pairwise identity | 61.23 |
| Shannon entropy | 0.79626 |
| G+C content | 0.62083 |
| Mean single sequence MFE | -28.21 |
| Consensus MFE | -16.84 |
| Energy contribution | -14.69 |
| Covariance contribution | -2.15 |
| Combinations/Pair | 2.18 |
| Mean z-score | -0.93 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.52 |
| SVM RNA-class probability | 0.992086 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
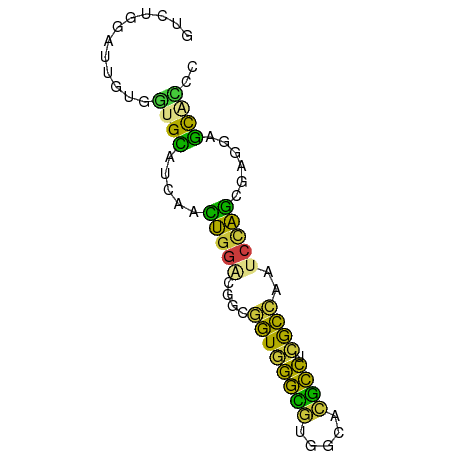
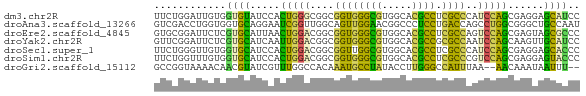
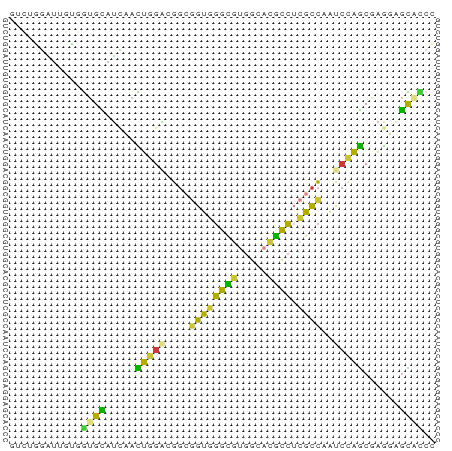
>dm3.chr2R 10546077 71 + 21146708 UUCUGGAUUGUGGUGUAUCCACUGGGCGGCGGUGGGCGUGGCACGCCUCGCCCAUCCAGCGAGGAGCAUCC ..((((((.((((.....)))).((((((.((((.((...)).))))))))))))))))............ ( -29.90, z-score = -0.51, R) >droAna3.scaffold_13266 5023569 71 - 19884421 GUCGACCUGGUGGUGCAGGAAUCGGUUGGCAGUUGGAACGGCCCUCCUGACCAGCCUGGCGGGCUGCCAAU (((((((.(((.........)))))))))).(((((..((((((.((.(......).)).))))))))))) ( -27.60, z-score = -0.02, R) >droEre2.scaffold_4845 7283902 71 - 22589142 GUGCGGAUUCUCGUGCAUUAACUGGACGGCGGUGGGCGUGGCACGCCUCGCCAGUCCAGCGAGUAGCGCCC ((((.....((((........((((((((((..(((((.....))))))))).))))))))))..)))).. ( -32.40, z-score = -1.33, R) >droYak2.chr2R 10477786 71 + 21139217 GUUCGGAUUCUCGUGCAUCAAUUGGACGGCGGUGGGCGUGGCACGCCGCGCCAAUCCAGCAAGUUGCAUCC ....((((.(.(((.((.....)).)))((((..(((((((....)))))))...)).)).....).)))) ( -26.80, z-score = -0.80, R) >droSec1.super_1 8065708 71 + 14215200 UUCUGGGUUGUGGUGCAUCCACUGGACGGCGGUUGGCGUGGCACGCCUCGCCCAUCCAGCGAGGAGCACCC ...........(((((.(((.(((((.(((((..((((.....)))))))))..)))))...)))))))). ( -32.70, z-score = -1.22, R) >droSim1.chr2R 9309000 71 + 19596830 UUCUGGUUUGUGGUGCAUCCACUGGACGGCGGUGGGCGUGGCACGCCUCGCCCGUCCAGCGAGGAGUACCC ...........(((((.(((.((((((((((..(((((.....))))))).))))))))...)))))))). ( -35.10, z-score = -1.92, R) >droGri2.scaffold_15112 4560740 67 - 5172618 GCCGGUAAAACAACGUAUCGUUUGGCCACAAAUGCCUAUACCUUGGGCCAUUUAA--AACAAAUAAUUU-- ...(((.((((........)))).)))..(((((((((.....)))).)))))..--............-- ( -13.00, z-score = -0.73, R) >consensus GUCUGGAUUGUGGUGCAUCAACUGGACGGCGGUGGGCGUGGCACGCCUCGCCAAUCCAGCGAGGAGCACCC ............((((.....(((((....((((((((.....)))).))))..)))))......)))).. (-16.84 = -14.69 + -2.15)

| Location | 10,546,077 – 10,546,148 |
|---|---|
| Length | 71 |
| Sequences | 7 |
| Columns | 71 |
| Reading direction | reverse |
| Mean pairwise identity | 61.23 |
| Shannon entropy | 0.79626 |
| G+C content | 0.62083 |
| Mean single sequence MFE | -27.00 |
| Consensus MFE | -15.25 |
| Energy contribution | -14.91 |
| Covariance contribution | -0.34 |
| Combinations/Pair | 1.80 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.02 |
| SVM RNA-class probability | 0.996978 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
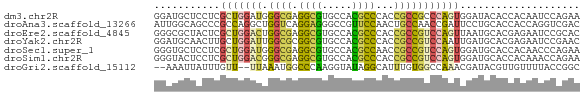
>dm3.chr2R 10546077 71 - 21146708 GGAUGCUCCUCGCUGGAUGGGCGAGGCGUGCCACGCCCACCGCCGCCCAGUGGAUACACCACAAUCCAGAA ((.....))...(((((((((((.((((.....))))......))))).((((.....)))).)))))).. ( -29.00, z-score = -1.74, R) >droAna3.scaffold_13266 5023569 71 + 19884421 AUUGGCAGCCCGCCAGGCUGGUCAGGAGGGCCGUUCCAACUGCCAACCGAUUCCUGCACCACCAGGUCGAC ..((((.....))))((.(((((((((((((.((....)).)))......)))))).))))))........ ( -23.80, z-score = -0.03, R) >droEre2.scaffold_4845 7283902 71 + 22589142 GGGCGCUACUCGCUGGACUGGCGAGGCGUGCCACGCCCACCGCCGUCCAGUUAAUGCACGAGAAUCCGCAC ..(((...((((((((((.((((.((((.....))))...))))))))))........))))....))).. ( -29.50, z-score = -1.59, R) >droYak2.chr2R 10477786 71 - 21139217 GGAUGCAACUUGCUGGAUUGGCGCGGCGUGCCACGCCCACCGCCGUCCAAUUGAUGCACGAGAAUCCGAAC ((((....(.((((.((((((.((((((((.......)).)))))))))))).).))).)...)))).... ( -25.00, z-score = -1.25, R) >droSec1.super_1 8065708 71 - 14215200 GGGUGCUCCUCGCUGGAUGGGCGAGGCGUGCCACGCCAACCGCCGUCCAGUGGAUGCACCACAACCCAGAA .((((((((..(((((((((.((.((((.....))))...)))))))))))))).)))))........... ( -36.60, z-score = -3.01, R) >droSim1.chr2R 9309000 71 - 19596830 GGGUACUCCUCGCUGGACGGGCGAGGCGUGCCACGCCCACCGCCGUCCAGUGGAUGCACCACAAACCAGAA (((((.(((..(((((((((.((.((((.....))))...))))))))))))))))).))........... ( -32.70, z-score = -2.24, R) >droGri2.scaffold_15112 4560740 67 + 5172618 --AAAUUAUUUGUU--UUAAAUGGCCCAAGGUAUAGGCAUUUGUGGCCAAACGAUACGUUGUUUUACCGGC --.......(((((--.....(((((((((((....)).)))).))))))))))...((((......)))) ( -12.40, z-score = 0.52, R) >consensus GGAUGCUACUCGCUGGAUGGGCGAGGCGUGCCACGCCCACCGCCGUCCAGUGGAUGCACCACAAUCCAGAC ...........(((((((.((((.((((.....))))...))))))))))).................... (-15.25 = -14.91 + -0.34)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:23:02 2011