
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 10,389,974 – 10,390,086 |
| Length | 112 |
| Max. P | 0.544627 |

| Location | 10,389,974 – 10,390,086 |
|---|---|
| Length | 112 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 67.19 |
| Shannon entropy | 0.65842 |
| G+C content | 0.40820 |
| Mean single sequence MFE | -23.91 |
| Consensus MFE | -10.41 |
| Energy contribution | -10.47 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.31 |
| Mean z-score | -0.86 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.544627 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 10389974 112 + 21146708 -GAAAAAUGACACAUUAAACGUUGUGACUCCUAAAAGGUAAGUGGCAUUUUCAAA-------CACCCACUUGUAGGCAGUCAAAAGGUAGACUGUUCACGGUCCAAAUAAUCGGAUGCAC -...........((((..((.((.(((((((((.....(((((((..........-------...))))))))))).))))).)).)).(((((....)))))..........))))... ( -24.32, z-score = -0.51, R) >droSim1.chr2R 9144456 119 + 19596830 -UGAAAAUUACACAUUAAACGUUUUGACUCCUAAAAGGAAAGCGGCAUUUUAAAAUAUUUCACACCCACCUGUAGGCAGUCAAAGGGUAGACUGUUCACGGUCCAGAUAAUCGGAUGCAC -..................(((((....(((.....))))))))(((......................((((..((((((........))))))..))))(((........)))))).. ( -24.10, z-score = -0.44, R) >droSec1.super_1 7900468 119 + 14215200 -UGAAAAUUACACAUUAAACGUUUUGACUCCUAAAAGGAAAGCGGCAUUUUCAAAUUGUUCACACCCACCUGUAGGCAGUCAAAAGGUAGACUGUUCACGGUCCAGAUAAUCGGAUGCAC -...........((((..((.((((((((((((..(((.....((((.........))))........))).)))).)))))))).)).(((((....)))))..........))))... ( -24.32, z-score = -0.23, R) >droYak2.chr2R 10315635 103 + 21139217 -UAAAAA-GAUAUAUUAAAUGCUUUGACUAUAAAAAGGAAAAG---------------CUCAGACCCACCUGUAGACAGUCAAAAGGUAGACUGUUGACGGUCCAAAUAAUUGGAUGCAC -......-............(((((..((......))..))))---------------)..........((((.(((((((........))))))).))))(((((....)))))..... ( -23.40, z-score = -2.56, R) >droEre2.scaffold_4845 7123352 118 - 22589142 -UAAAAAUGACACAUUAAACGUUUUGACUCUUAAAAGAAAAAGGGAAUUUUAAUA-AGUUCAGACCAACCUGUAGACAGUCAAAUGGAAGACUGUUCACGGUCCAAAUAAUCGGAUGCAC -...........((((......((((.(((((........)))))..........-......((((.....((..((((((........))))))..))))))))))......))))... ( -20.90, z-score = -0.45, R) >droAna3.scaffold_13266 14715178 100 + 19884421 ----------AGGAUCAAG---UUUUAGUUUUAAAGAACAAGAGCUAUAUU-------UUUAGACCCACCUGCUGGCAAUCGAAGGGCAGACUGUUGACAGUCCAGAUAAUAGGAUGCAC ----------.((.((.((---((((.((((....)))).)))))).((..-------..)))).)).(((..((.....)).)))(((..((((((.(......).))))))..))).. ( -18.90, z-score = 0.22, R) >dp4.chr3 5017511 104 + 19779522 CAAAAAGCUGAAGAUUAAC--UUUCGGCUCAAAUGAUACUGUUGGGG--------------GAACACACCUGGAAGCAGUCGAAGGGCAGACUAUUCACUGUCCAUAUAAUCGGAUGGAC .....((((((((......--))))))))((..((((((((((.(((--------------.......)))...))))))....((((((........)))))).....))))..))... ( -27.80, z-score = -2.00, R) >droPer1.super_2 5216838 104 + 9036312 CAAAGAGCUGGAGAUUAAA--UUUCGGCCCAAAUUAUGCUGUUGGGG--------------GAACACACCUGGAAGCAGUCGAAGGGCAGACUAUUCACUGUCCAUAUAAUCGGAUGGAC ......(((((((......--)))))))(((.....((((.(..((.--------------.......))..).))))......((((((........))))))...........))).. ( -27.50, z-score = -0.90, R) >consensus _UAAAAAUUACACAUUAAACGUUUUGACUCCUAAAAGGAAAGUGGCAUUUU_______UUCACACCCACCUGUAGGCAGUCAAAGGGUAGACUGUUCACGGUCCAAAUAAUCGGAUGCAC .....................................................................(((..(((((((........)))))))..)))(((........)))..... (-10.41 = -10.47 + 0.06)

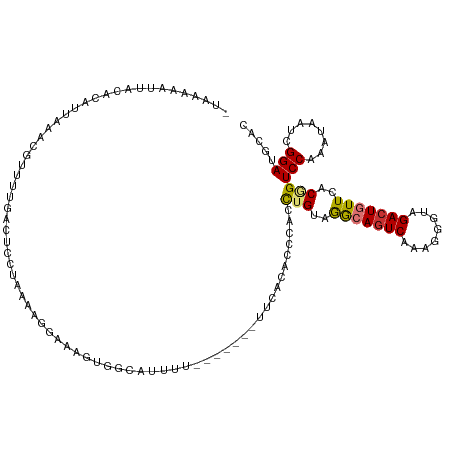

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:22:40 2011