
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 10,064,638 – 10,064,744 |
| Length | 106 |
| Max. P | 0.871017 |

| Location | 10,064,638 – 10,064,744 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 84.97 |
| Shannon entropy | 0.29268 |
| G+C content | 0.57407 |
| Mean single sequence MFE | -37.77 |
| Consensus MFE | -29.73 |
| Energy contribution | -30.98 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.775927 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
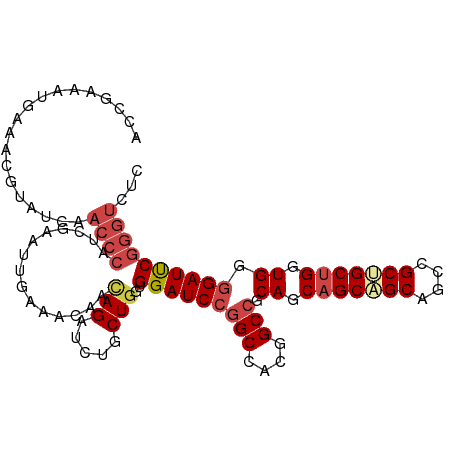
>dm3.chr2R 10064638 106 + 21146708 ACCGAAAUGAAACGUAUCAACCCAUCGAAUUGAAAAAACAGAUCUGCUAGUGGAUCCGGCCACGGCCGCAGCAGCUGCAGCCGCCGCUGGUGGGGAUUCGGGUCUC .(((((.(((......))).((((((..............((((((....)))))).(((..((((.((((...)))).))))..)))))))))..)))))..... ( -37.20, z-score = -1.30, R) >droSim1.chr2R 8834145 106 + 19596830 ACCGAAAUGAAACGUAUCAACCCAUCGAAUUGAAACAACAGAUCUGCUGGUGGAUCCGGCCACGGCCGCAGCAGCAGCAGCCGCUGCUGGUGGGGAUUCGGGUCUC .(((((.(((......))).((((((((.(((......))).)).((((.(((..(((....))))))))))((((((....))))))))))))..)))))..... ( -39.40, z-score = -1.55, R) >droSec1.super_1 7572303 106 + 14215200 ACCGAAAUGAAACGUAUCAACCCAUCGAAUUGAAACAACAGAUCUGCUGGUGGAUCCGGCCACGGCCGCAGCAGCAGCAGCCGCUGCUGGUGGGGAUUCGGGUCUC .(((((.(((......))).((((((((.(((......))).)).((((.(((..(((....))))))))))((((((....))))))))))))..)))))..... ( -39.40, z-score = -1.55, R) >droYak2.chr2R 10000741 106 + 21139217 ACCGAAAUGAAACGUAUCAACCCAUCGAAUUGAAACAACAGAUCUGCUGGUGGAUCCGGCCACGGCCGCAGCAGCUGCAGCCGCUGCUGGUGGGGAUUCGGGUCUC .(((((.(((......))).(((((((.............((((((....)))))).(((....)))(((((.((....)).))))))))))))..)))))..... ( -39.20, z-score = -1.45, R) >droEre2.scaffold_4845 6815727 106 - 22589142 ACCAAAAUGAAACGUACCAACCCAUCGAAUUGAAACAACAGAUGUGCUGGUGGAUCCGGCCACGGCCGCAGCAGCAGCAGCCGCUGCUGGUGGGGAUUCGGGUCUC .((............((((.(.((((...(((...)))..)))).).))))(((((((((....))).((.(((((((....))))))).)).))))))))..... ( -40.00, z-score = -1.59, R) >droAna3.scaffold_13266 14027725 85 - 19884421 ---------------ACAAA--AAUGGAACCACAA-AUUAGAUCUUCUGGUGGAUCAGGCCACGGCAGCAGCAGCUGCAGCCGCGGCGGCUGGGGAUCCGACA--- ---------------.....--..((((.((.((.-....(((((......)))))..(((.((((.((((...)))).)))).)))...)).)).))))...--- ( -31.40, z-score = -1.73, R) >consensus ACCGAAAUGAAACGUAUCAACCCAUCGAAUUGAAACAACAGAUCUGCUGGUGGAUCCGGCCACGGCCGCAGCAGCAGCAGCCGCUGCUGGUGGGGAUUCGGGUCUC ...................((((...............(((.....)))..(((((((((....))).((.(((((((....))))))).)).))))))))))... (-29.73 = -30.98 + 1.25)


| Location | 10,064,638 – 10,064,744 |
|---|---|
| Length | 106 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 84.97 |
| Shannon entropy | 0.29268 |
| G+C content | 0.57407 |
| Mean single sequence MFE | -39.20 |
| Consensus MFE | -28.78 |
| Energy contribution | -30.67 |
| Covariance contribution | 1.89 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.871017 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
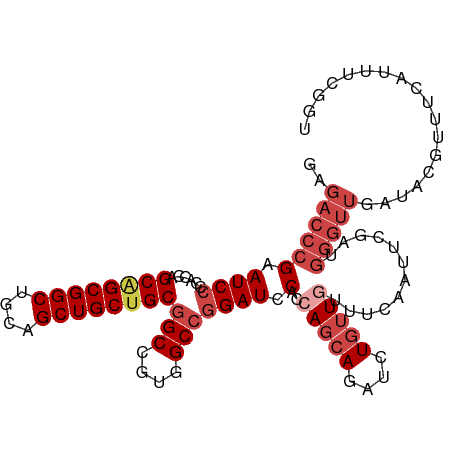
>dm3.chr2R 10064638 106 - 21146708 GAGACCCGAAUCCCCACCAGCGGCGGCUGCAGCUGCUGCGGCCGUGGCCGGAUCCACUAGCAGAUCUGUUUUUUCAAUUCGAUGGGUUGAUACGUUUCAUUUCGGU .....(((((..((((.(.((.((((((((((...)))))))))).))((((((........))))))............).)))).(((......))).))))). ( -40.10, z-score = -1.85, R) >droSim1.chr2R 8834145 106 - 19596830 GAGACCCGAAUCCCCACCAGCAGCGGCUGCUGCUGCUGCGGCCGUGGCCGGAUCCACCAGCAGAUCUGUUGUUUCAAUUCGAUGGGUUGAUACGUUUCAUUUCGGU .....(((((..((((((.((((((((....))))))))(((....)))))......(((((....)))))...........)))).(((......))).))))). ( -39.70, z-score = -1.61, R) >droSec1.super_1 7572303 106 - 14215200 GAGACCCGAAUCCCCACCAGCAGCGGCUGCUGCUGCUGCGGCCGUGGCCGGAUCCACCAGCAGAUCUGUUGUUUCAAUUCGAUGGGUUGAUACGUUUCAUUUCGGU .....(((((..((((((.((((((((....))))))))(((....)))))......(((((....)))))...........)))).(((......))).))))). ( -39.70, z-score = -1.61, R) >droYak2.chr2R 10000741 106 - 21139217 GAGACCCGAAUCCCCACCAGCAGCGGCUGCAGCUGCUGCGGCCGUGGCCGGAUCCACCAGCAGAUCUGUUGUUUCAAUUCGAUGGGUUGAUACGUUUCAUUUCGGU .....(((((..((((((.((((((((....))))))))(((....)))))......(((((....)))))...........)))).(((......))).))))). ( -39.40, z-score = -1.59, R) >droEre2.scaffold_4845 6815727 106 + 22589142 GAGACCCGAAUCCCCACCAGCAGCGGCUGCUGCUGCUGCGGCCGUGGCCGGAUCCACCAGCACAUCUGUUGUUUCAAUUCGAUGGGUUGGUACGUUUCAUUUUGGU (((((..(.((((......((((((((....))))))))(((....))))))).)((((((.((((.((((...))))..)))).))))))..)))))........ ( -43.70, z-score = -2.53, R) >droAna3.scaffold_13266 14027725 85 + 19884421 ---UGUCGGAUCCCCAGCCGCCGCGGCUGCAGCUGCUGCUGCCGUGGCCUGAUCCACCAGAAGAUCUAAU-UUGUGGUUCCAUU--UUUGU--------------- ---....(((.((.(((..((((((((.((((...)))).))))))))..((((........))))....-))).)).)))...--.....--------------- ( -32.60, z-score = -2.48, R) >consensus GAGACCCGAAUCCCCACCAGCAGCGGCUGCAGCUGCUGCGGCCGUGGCCGGAUCCACCAGCAGAUCUGUUGUUUCAAUUCGAUGGGUUGAUACGUUUCAUUUCGGU ..((((((.((((......((((((((....))))))))(((....))))))).)..(((((....)))))............))))).................. (-28.78 = -30.67 + 1.89)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:21:59 2011