
| Sequence ID | dm3.chr2LHet |
|---|---|
| Location | 121,787 – 121,907 |
| Length | 120 |
| Max. P | 0.996050 |

| Location | 121,787 – 121,907 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 61.94 |
| Shannon entropy | 0.63417 |
| G+C content | 0.38406 |
| Mean single sequence MFE | -24.73 |
| Consensus MFE | -14.96 |
| Energy contribution | -15.65 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.88 |
| SVM RNA-class probability | 0.996050 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
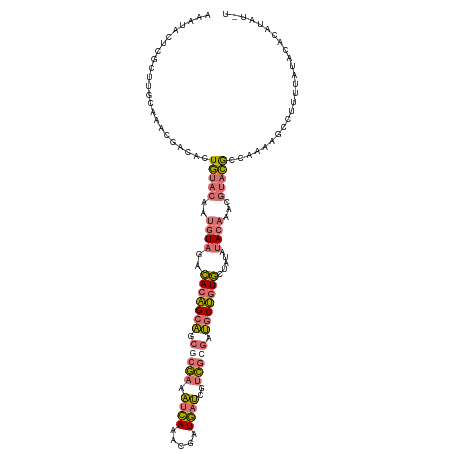
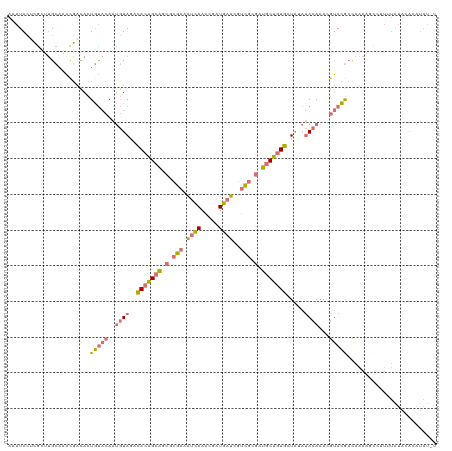
>dm3.chr2LHet 121787 120 + 368872 AAAUACUCGCUUGCAAACGACACUGUACAAUGUAAUCACAGCAGCGCGAAAUCAAACGAUGAUCGUCGCGAUGCUGUGCUAAAUACAAACGUACGCCAAAAGCCUUUUAUACACAUAUUU ........((((...........(((((..((((..(((((((.(((((.((((.....))))..))))).))))))).....))))...)))))....))))................. ( -27.26, z-score = -1.47, R) >droYak2.chrU 27704695 120 - 28119190 AAAUACUCGCUUGCAAACGACACUAUACAAUGUAGACACAGCAACACGAAAUCAAACGAUGAUCGUCGCGAUGCUGUGCUAUAUACAAACGUAUGCCAAAAGCCUUUUAUACACAUAUGU ........(((((((.(((..........((((((.(((((((.(.(((.((((.....))))..))).).))))))))))))).....))).)))...))))................. ( -27.86, z-score = -2.42, R) >droEre2.scaffold_4684 28488 120 + 132001 AAAUACCCACGUGCAAACGGCACUGUACAAUGUAGCCACAGCAGCGCGAAAUCAAACGAUGAUCGUCGCGAUGCUGUGCUGUAUACAAAAGUACGCCAAAUUUUUUUUAUAUACAUAUAC ..........((((.....)))).((((..((((..(((((((.(((((.((((.....))))..))))).))))))).....))))...)))).......................... ( -32.40, z-score = -2.19, R) >anoGam1.chrU 13855738 97 - 59568033 UAAUAAUAGCUGGCGAUUU-UAAUGCACAUCAUACAUACUGGGGAGAUGAUAGAAUCGAUCAAAAAGGAAACACAAUAAUAGA-AGAAAUAGACAAUUC--------------------- ..........(((((((((-((...((..((...(.....)..))..)).)))))))).)))....(....)...........-...............--------------------- ( -11.40, z-score = -0.86, R) >consensus AAAUACUCGCUUGCAAACGACACUGUACAAUGUAGACACAGCAGCGCGAAAUCAAACGAUGAUCGUCGCGAUGCUGUGCUAUAUACAAACGUACGCCAAAAGCCUUUUAUACACAUAU_U .......................(((((..((((..(((((((.(((((.((((.....))))..))))).))))))).....))))...)))))......................... (-14.96 = -15.65 + 0.69)


| Location | 121,787 – 121,907 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 61.94 |
| Shannon entropy | 0.63417 |
| G+C content | 0.38406 |
| Mean single sequence MFE | -29.95 |
| Consensus MFE | -22.11 |
| Energy contribution | -22.68 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.25 |
| SVM RNA-class probability | 0.986738 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
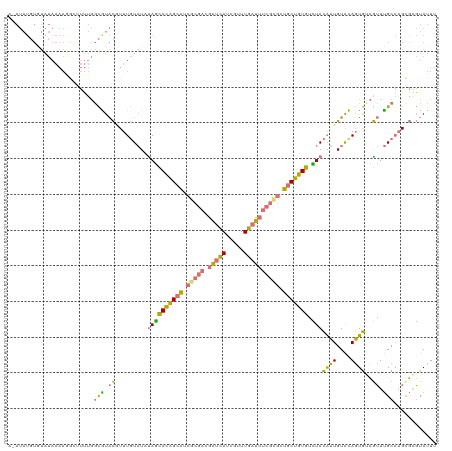
>dm3.chr2LHet 121787 120 - 368872 AAAUAUGUGUAUAAAAGGCUUUUGGCGUACGUUUGUAUUUAGCACAGCAUCGCGACGAUCAUCGUUUGAUUUCGCGCUGCUGUGAUUACAUUGUACAGUGUCGUUUGCAAGCGAGUAUUU ((((((...........((.....))...((((((((.....(((((((.(((((.(((((.....)))))))))).)))))))...(((((....)))))....)))))))).)))))) ( -37.30, z-score = -1.66, R) >droYak2.chrU 27704695 120 + 28119190 ACAUAUGUGUAUAAAAGGCUUUUGGCAUACGUUUGUAUAUAGCACAGCAUCGCGACGAUCAUCGUUUGAUUUCGUGUUGCUGUGUCUACAUUGUAUAGUGUCGUUUGCAAGCGAGUAUUU .................((((...(((.(((.(((((((((((((((((.(((((.(((((.....)))))))))).))))))).)))...)))))))...))).)))))))........ ( -37.40, z-score = -2.17, R) >droEre2.scaffold_4684 28488 120 - 132001 GUAUAUGUAUAUAAAAAAAAUUUGGCGUACUUUUGUAUACAGCACAGCAUCGCGACGAUCAUCGUUUGAUUUCGCGCUGCUGUGGCUACAUUGUACAGUGCCGUUUGCACGUGGGUAUUU ((((((((((((((((..............))))))))))).(((((((.(((((.(((((.....)))))))))).))))))).......))))).((((.....)))).......... ( -39.64, z-score = -2.51, R) >anoGam1.chrU 13855738 97 + 59568033 ---------------------GAAUUGUCUAUUUCU-UCUAUUAUUGUGUUUCCUUUUUGAUCGAUUCUAUCAUCUCCCCAGUAUGUAUGAUGUGCAUUA-AAAUCGCCAGCUAUUAUUA ---------------------((((((((.......-......................)).))))))(((((((..........).)))))).......-................... ( -5.45, z-score = 1.96, R) >consensus A_AUAUGUGUAUAAAAGGCUUUUGGCGUACGUUUGUAUAUAGCACAGCAUCGCGACGAUCAUCGUUUGAUUUCGCGCUGCUGUGUCUACAUUGUACAGUGUCGUUUGCAAGCGAGUAUUU ........................(((.(((.(((((((((((((((((.(((((.(((((.....)))))))))).))))))).)))...)))))))...))).)))............ (-22.11 = -22.68 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:04:45 2011