
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 9,318,485 – 9,318,559 |
| Length | 74 |
| Max. P | 0.595472 |

| Location | 9,318,485 – 9,318,559 |
|---|---|
| Length | 74 |
| Sequences | 12 |
| Columns | 74 |
| Reading direction | reverse |
| Mean pairwise identity | 79.92 |
| Shannon entropy | 0.48213 |
| G+C content | 0.56962 |
| Mean single sequence MFE | -19.63 |
| Consensus MFE | -14.49 |
| Energy contribution | -13.57 |
| Covariance contribution | -0.93 |
| Combinations/Pair | 1.82 |
| Mean z-score | -0.62 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.595472 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
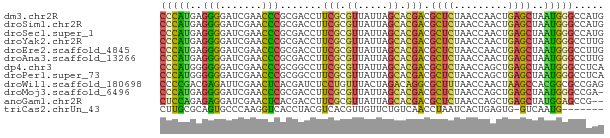
>dm3.chr2R 9318485 74 - 21146708 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCAUG (((((..(((.......)))(((.....)))..............((((.........))))..)))))..... ( -20.00, z-score = -0.97, R) >droSim1.chr2R 7772076 74 - 19596830 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCAUG (((((..(((.......)))(((.....)))..............((((.........))))..)))))..... ( -20.00, z-score = -0.97, R) >droSec1.super_1 6836815 74 - 14215200 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCAUG (((((..(((.......)))(((.....)))..............((((.........))))..)))))..... ( -20.00, z-score = -0.97, R) >droYak2.chr2R 13497887 74 + 21139217 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCUUG (((((..(((.......)))(((.....)))..............((((.........))))..)))))..... ( -20.00, z-score = -0.98, R) >droEre2.scaffold_4845 6089674 74 + 22589142 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCUUG (((((..(((.......)))(((.....)))..............((((.........))))..)))))..... ( -20.00, z-score = -0.98, R) >droAna3.scaffold_13266 10206247 74 + 19884421 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCUUG (((((..(((.......)))(((.....)))..............((((.........))))..)))))..... ( -20.00, z-score = -0.98, R) >dp4.chr3 1802855 74 + 19779522 CCCAUGGGGGGAUCGAACCCGCGGCCUUCGCGUUAUUAGCACGACGCUCUAACCAGCUGAGCUAAUGGGCCUCA .....(.(((.......))).)((((((((.((.....)).))).((((.........))))....)))))... ( -23.20, z-score = 0.19, R) >droPer1.super_73 79876 74 - 267513 CCCAUGGGGGGAUCGAACCCGCGGCCUUCGCGUUAUUAGCACGACGCUCUAACCAGCUGAGCUAAUGGGCCUCA .....(.(((.......))).)((((((((.((.....)).))).((((.........))))....)))))... ( -23.20, z-score = 0.19, R) >droWil1.scaffold_180698 9356595 74 + 11422946 CCCCGACGAGAUUCGAACUCACGAUCUCCUGUUUACUAGACAGGCGCUUUAACCAACUAAGCCACGGCGCCGAG .......(((((.((......))))))).((((.....))))((((((.................))))))... ( -16.33, z-score = -1.22, R) >droMoj3.scaffold_6496 13658313 73 - 26866924 CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAGCUGAGCUAAUGGGCCGA- (((....)))..(((..((((((.....))))..((((((.((..(((......))))).))))))))..)))- ( -21.40, z-score = -0.57, R) >anoGam1.chr2R 13943283 72 + 62725911 CUCCAGAGAGGAUCGAACUCACGACCUUCGCGUUAUUAGCACGACGCUCUAACCAGCUGAGCUAUGGAGCCG-- (((((..((((.(((......))).))))((.......)).....((((.........))))..)))))...-- ( -19.00, z-score = -0.79, R) >triCas2.chrUn_43 97726 66 + 275627 CUUGCGCAGUGCCCAAGGUCACCUACGUCACGUUGUUCUGUCAACCUAAUCACUGAGUG-GUCAAUG------- .(..(.(((((....((((.((..(((......)))...))..))))...))))).)..-)......------- ( -12.40, z-score = 0.56, R) >consensus CCCAUGAGGGGAUCGAACCCGCGACCUUCGCGUUAUUAGCACGACGCUCUAACCAACUGAGCUAAUGGGCCUUG (((((..(((.......))).......(((.((.....)).))).((((.........))))..)))))..... (-14.49 = -13.57 + -0.93)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:19:59 2011