
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 9,216,905 – 9,217,000 |
| Length | 95 |
| Max. P | 0.999430 |

| Location | 9,216,905 – 9,217,000 |
|---|---|
| Length | 95 |
| Sequences | 14 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 88.75 |
| Shannon entropy | 0.26743 |
| G+C content | 0.50434 |
| Mean single sequence MFE | -39.88 |
| Consensus MFE | -33.18 |
| Energy contribution | -33.75 |
| Covariance contribution | 0.58 |
| Combinations/Pair | 1.04 |
| Mean z-score | -4.68 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.88 |
| SVM RNA-class probability | 0.999430 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
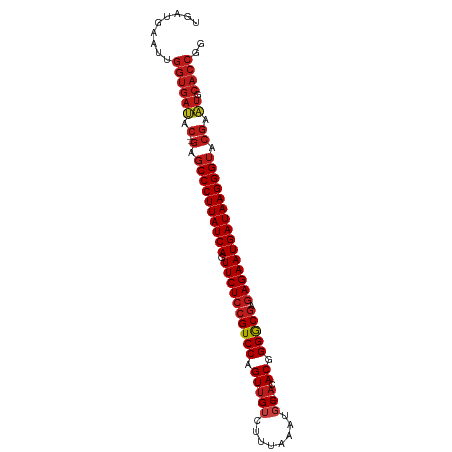
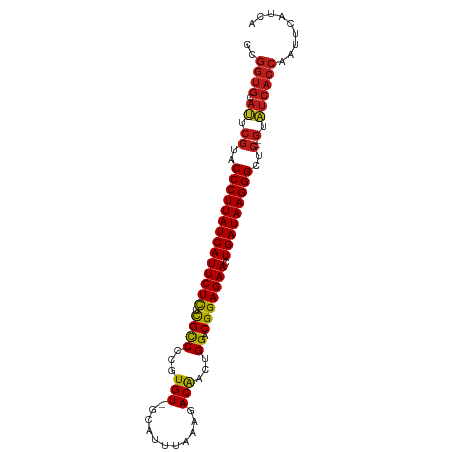
>dm3.chr2R 9216905 95 + 21146708 UGAUGAAUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUAAGUGC-ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -5.01, R) >apiMel3.Group13 1186510 83 + 8929068 -GAUGAUUCUGUGACCCUCGGGCCCUUAUCAGUUCUCCGUCCAGUCGUCUAACAAUUCUACACCGGACAGGAGAAUGAUAAGGG-------------- -.............((....))(((((((((.((((((((((.((.((...........)))).)))).)))))))))))))))-------------- ( -28.80, z-score = -2.47, R) >droSim1.chr2R 7673839 95 + 19596830 UGAUGAAUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUAAAUGC-ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -5.27, R) >droSec1.super_1 6740016 95 + 14215200 UGAUGAAUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUAAAUGC-ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -5.27, R) >droYak2.chr2R 13396856 94 - 21139217 UGAUGAAUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUCAA-UGC-ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((......-.))-).)).))))).))))))))))))))).)).)).))))). ( -40.30, z-score = -4.79, R) >droEre2.scaffold_4845 5990942 95 - 22589142 UGACGAAUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUAAAUGC-ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -5.17, R) >droAna3.scaffold_13266 16524578 95 + 19884421 UGAUGAAUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUCAUUAUGC-ACACGGGGCGCGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -4.57, R) >dp4.chr3 19082706 95 - 19779522 UGAUGAGUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUUAAUGC-ACACGGGGCGCGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -4.68, R) >droPer1.super_2 8173054 95 - 9036312 UGAUGAGUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUUAAUGC-ACACGGGGCGCGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -4.68, R) >droWil1.scaffold_181009 2073161 95 + 3585778 UGUUGAGUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUAGAUAUGC-ACACGGGGCGCGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -4.39, R) >droVir3.scaffold_12875 20368191 95 - 20611582 AGUUGACUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUAGAUAUGC-ACACGGGGCGCGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((........))-).)).))))).))))))))))))))).)).)).))))). ( -40.60, z-score = -4.44, R) >droMoj3.scaffold_6496 5255796 95 + 26866924 AGUUGACUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUAUAGAUAUGC-ACACGGGGCGCGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((.(((((((....))))-).)).))))).))))))))))))))).)).)).))))). ( -43.20, z-score = -5.60, R) >droGri2.scaffold_15245 10719552 95 + 18325388 UGAUGAGUUGGUGAUAC--GAGCCCUUAUCAGUUCUCCGUCCAGUUGUAUAGUUAUCC-ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG ........(((((((.(--(.((((((((((.((((((((((...(((..........-)))..))))).))))))))))))))).)).)).))))). ( -38.90, z-score = -4.52, R) >anoGam1.chr3R 9950677 91 - 53272125 GGGUAAA--GGUGACCC--GGGCCCUUAUCAGUUCUCCGUCCAGUUGU-UAGAAA-GC-ACACGGGGCGAAAGAAUGAUAAGGGUUCGAGUGCACCGG .......--((((((.(--((((((((((((.((((.(((((.(((((-(....)-))-).)).)))))..))))))))))))))))).)).)))).. ( -41.70, z-score = -4.67, R) >consensus UGAUGAAUUGGUGAUAC__GAGCCCUUAUCAGUUCUCCGUCCAGUUGUCUUUAAAUGC_ACACGGGGCGAGAGAAUGAUAAGGGUACGAAUGCACCGG .........((((........((((((((((.((((((((((.((.((........)).))...))))).))))))))))))))).......)))).. (-33.18 = -33.75 + 0.58)


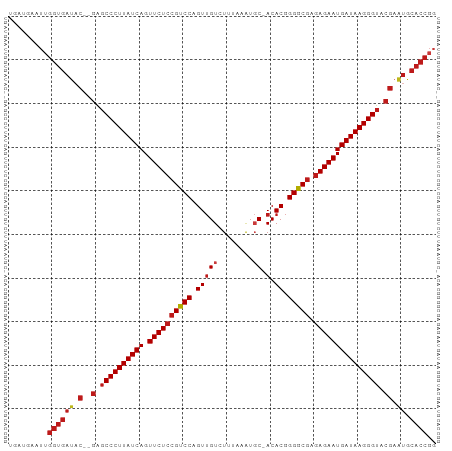
| Location | 9,216,905 – 9,217,000 |
|---|---|
| Length | 95 |
| Sequences | 14 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 88.75 |
| Shannon entropy | 0.26743 |
| G+C content | 0.50434 |
| Mean single sequence MFE | -34.78 |
| Consensus MFE | -27.14 |
| Energy contribution | -27.30 |
| Covariance contribution | 0.15 |
| Combinations/Pair | 1.08 |
| Mean z-score | -4.33 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.82 |
| SVM RNA-class probability | 0.999353 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 9216905 95 - 21146708 CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU-GCACUUAAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAUUCAUCA ..((((.((.((..((((((((((((((.((((...(((-..........)))...)).))))))).)))))))))..)--).))))))......... ( -33.70, z-score = -4.42, R) >apiMel3.Group13 1186510 83 - 8929068 --------------CCCUUAUCAUUCUCCUGUCCGGUGUAGAAUUGUUAGACGACUGGACGGAGAACUGAUAAGGGCCCGAGGGUCACAGAAUCAUC- --------------(((((((((((((((.(((((((((...........)).))))))))))))).)))))))))((....)).............- ( -33.40, z-score = -3.37, R) >droSim1.chr2R 7673839 95 - 19596830 CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU-GCAUUUAAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAUUCAUCA ..((((.((.((..((((((((((((((.((((...(((-..........)))...)).))))))).)))))))))..)--).))))))......... ( -33.70, z-score = -4.46, R) >droSec1.super_1 6740016 95 - 14215200 CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU-GCAUUUAAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAUUCAUCA ..((((.((.((..((((((((((((((.((((...(((-..........)))...)).))))))).)))))))))..)--).))))))......... ( -33.70, z-score = -4.46, R) >droYak2.chr2R 13396856 94 + 21139217 CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU-GCA-UUGAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAUUCAUCA ..((((.((.((..((((((((((((((.((((...(((-.(.-.....))))...)).))))))).)))))))))..)--).))))))......... ( -34.20, z-score = -3.94, R) >droEre2.scaffold_4845 5990942 95 + 22589142 CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU-GCAUUUAAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAUUCGUCA ..((((.((.((..((((((((((((((.((((...(((-..........)))...)).))))))).)))))))))..)--).))))))......... ( -33.70, z-score = -4.07, R) >droAna3.scaffold_13266 16524578 95 - 19884421 CCGGUGCAUUCGUACCCUUAUCAUUCUCGCGCCCCGUGU-GCAUAAUGAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAUUCAUCA ..((((.((.((..((((((((((((((.((((...(((-.(.....)..)))...)).))))))).)))))))))..)--).))))))......... ( -34.10, z-score = -3.57, R) >dp4.chr3 19082706 95 + 19779522 CCGGUGCAUUCGUACCCUUAUCAUUCUCGCGCCCCGUGU-GCAUUAAAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAACUCAUCA ..((((.((.((..((((((((((((((.((((...(((-..........)))...)).))))))).)))))))))..)--).))))))......... ( -33.70, z-score = -4.06, R) >droPer1.super_2 8173054 95 + 9036312 CCGGUGCAUUCGUACCCUUAUCAUUCUCGCGCCCCGUGU-GCAUUAAAAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAACUCAUCA ..((((.((.((..((((((((((((((.((((...(((-..........)))...)).))))))).)))))))))..)--).))))))......... ( -33.70, z-score = -4.06, R) >droWil1.scaffold_181009 2073161 95 - 3585778 CCGGUGCAUUCGUACCCUUAUCAUUCUCGCGCCCCGUGU-GCAUAUCUAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAACUCAACA ..((((.((.((..((((((((((((((((((.....))-))...(((((....)))))..))))).)))))))))..)--).))))))......... ( -37.00, z-score = -5.01, R) >droVir3.scaffold_12875 20368191 95 + 20611582 CCGGUGCAUUCGUACCCUUAUCAUUCUCGCGCCCCGUGU-GCAUAUCUAGACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAGUCAACU ..((((.((.((..((((((((((((((((((.....))-))...(((((....)))))..))))).)))))))))..)--).))))))......... ( -37.00, z-score = -4.63, R) >droMoj3.scaffold_6496 5255796 95 - 26866924 CCGGUGCAUUCGUACCCUUAUCAUUCUCGCGCCCCGUGU-GCAUAUCUAUACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAAGUCAACU ..((((.((.((..((((((((((((((.((((...(((-(........))))...)).))))))).)))))))))..)--).))))))......... ( -35.00, z-score = -4.39, R) >droGri2.scaffold_15245 10719552 95 - 18325388 CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU-GGAUAACUAUACAACUGGACGGAGAACUGAUAAGGGCUC--GUAUCACCAACUCAUCA ..((((.((.((..((((((((((((((.((((..((((-((....))))))....)).))))))).)))))))))..)--).))))))......... ( -37.00, z-score = -5.29, R) >anoGam1.chr3R 9950677 91 + 53272125 CCGGUGCACUCGAACCCUUAUCAUUCUUUCGCCCCGUGU-GC-UUUCUA-ACAACUGGACGGAGAACUGAUAAGGGCCC--GGGUCACC--UUUACCC ..((((.(((((..((((((((((((((.((((...(((-..-......-)))...)).))))))).)))))))))..)--))))))))--....... ( -37.00, z-score = -4.84, R) >consensus CCGGUGCAUUCGUACCCUUAUCAUUCUCUCGCCCCGUGU_GCAUUUAAAGACAACUGGACGGAGAACUGAUAAGGGCUC__GUAUCACCAAUUCAUCA ..((((........((((((((((((((.((.((.((................)).)).))))))).))))))))).........))))......... (-27.14 = -27.30 + 0.15)


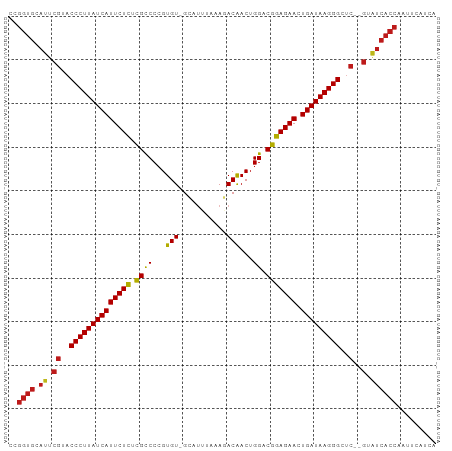
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:19:47 2011