
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 9,182,790 – 9,182,886 |
| Length | 96 |
| Max. P | 0.799931 |

| Location | 9,182,790 – 9,182,886 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 121 |
| Reading direction | forward |
| Mean pairwise identity | 65.03 |
| Shannon entropy | 0.62140 |
| G+C content | 0.59755 |
| Mean single sequence MFE | -34.73 |
| Consensus MFE | -16.17 |
| Energy contribution | -16.46 |
| Covariance contribution | 0.30 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.73 |
| SVM RNA-class probability | 0.799931 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 9182790 96 + 21146708 -----GCUGGCACAAGGAAUUCC---UCCUAGGCAGCCUGUCGCAUAUCCUGCGGCGAUGCAGUGACUUUUCCUGACUUCCUUCUGGCCUGCCUCGUGGCAAAG----------------- -----..((.(((.((((.....---))))((((((((.((((((.....))))))((..(((.((....)))))...)).....)).)))))).))).))...----------------- ( -33.10, z-score = -1.31, R) >droEre2.scaffold_4845 5957369 94 - 22589142 -----GCUGGCACAAGGAAUUCC---UCCUAGGCAGCCCGCCGCAUAUCCUGCGGCGAUGCAGUGAGUUCCCC--GCUUCCUUCUGGCCUGCCUCCUGCCAAAG----------------- -----..(((((..((((.....---))))(((((((((((((((.....)))))))..(.((..(((.....--)))..)).).)).))))))..)))))...----------------- ( -37.20, z-score = -2.73, R) >droYak2.chr2R 13362327 96 - 21139217 -----GCUGGCACAAGGAAUUCC---UCCUAGGCAACCCGCCGCAUAUCCUGCGGCGAUGCAGUGACUUUUCCUCGCCCCCUUCUGGCCCGCCUCCUGGCAAAG----------------- -----((((((.(.((((.....---)))).)))....(((((((.....)))))))...))))...........(((.......)))..(((....)))....----------------- ( -30.90, z-score = -1.21, R) >droSec1.super_1 6705510 96 + 14215200 -----GCUGGCACAAGGAAUUCC---UCCUAGGCAGCCUGCCGCAUAUCCUGCGGCGAUGCAGUGACUCUUCCUGACUUCCUUCUGGCCUGCCUCGUGGCAAAG----------------- -----..((.(((.((((.....---))))((((((((.((((((.....))))))((..(((.((....)))))...)).....)).)))))).))).))...----------------- ( -35.80, z-score = -1.91, R) >droSim1.chr2R 7641286 96 + 19596830 -----GCUGGCACAAGGAAUUCC---UCCUAGGCAGCCUGCCGCAUAUCCUGCGGCGAUGCAGUGACUCUUCCUGACUUCCUUCUGGCCUGCCUCGUGGCAAAG----------------- -----..((.(((.((((.....---))))((((((((.((((((.....))))))((..(((.((....)))))...)).....)).)))))).))).))...----------------- ( -35.80, z-score = -1.91, R) >dp4.chr3 19045000 117 - 19779522 CUCUUGCUGCUGGGUGGAAACUC---UUCUAAGCAACUUGCCGCAUAUCCUGAGGCGAUGCAGUGGCUCGGGU-GCCGGUCCUGGUGCUGCUCCUAGGUCUUGGUCCUUUUCCCGGCACUA .......(((((((.(((.....---.((((.(((..(((((.((.....)).))))))))((..(((((((.-......))))).))..))..))))......)))....)))))))... ( -39.80, z-score = 0.11, R) >droPer1.super_2 8133742 117 - 9036312 CUCUUGCUGCUGGGUGGAAACUC---UUCUAAGCAACUUGCCGCAUAUCCUGAGGCGAUGCAGUGGCUCGGGU-GCCGGUCCUGGUGCUGCUCCUAGGUCUUGGUCCUUUUCCCGGCACUA .......(((((((.(((.....---.((((.(((..(((((.((.....)).))))))))((..(((((((.-......))))).))..))..))))......)))....)))))))... ( -39.80, z-score = 0.11, R) >droAna3.scaffold_13266 16490955 81 + 19884421 -----UCUCGCACAAGGAAUUCCGCCUCCUAGGCAGCCCGUCGCAUAUCCUGCGGCGUUGCAGUCCUUUUUGG------CCCUGGGACCAAG----------------------------- -----........(((((.....(((.....))).((.(((((((.....)))))))..))..))))).((((------((...)).)))).----------------------------- ( -25.40, z-score = -0.51, R) >consensus _____GCUGGCACAAGGAAUUCC___UCCUAGGCAGCCUGCCGCAUAUCCUGCGGCGAUGCAGUGACUCUUCC_GACUUCCUUCUGGCCUGCCUCGUGGCAAAG_________________ .....(((((....((((........))))..(((...(((((((.....))))))).)))......................)))))................................. (-16.17 = -16.46 + 0.30)

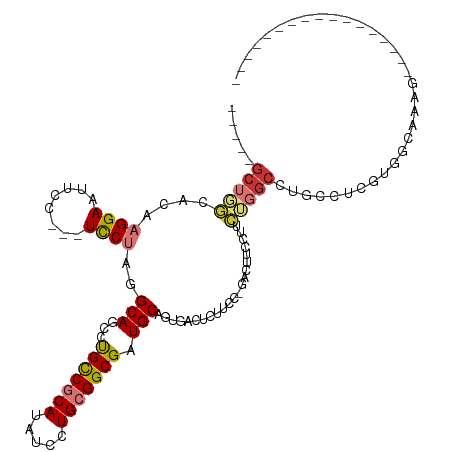
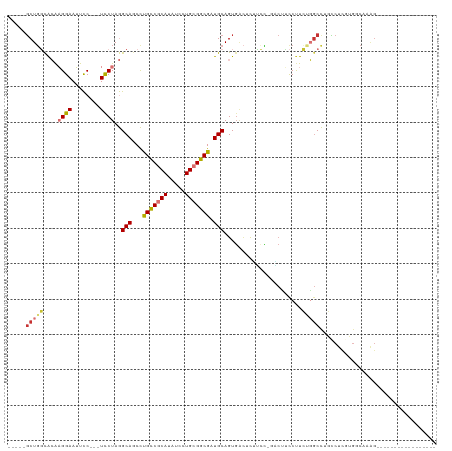
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:19:35 2011