
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 2,721,872 – 2,721,974 |
| Length | 102 |
| Max. P | 0.987271 |

| Location | 2,721,872 – 2,721,974 |
|---|---|
| Length | 102 |
| Sequences | 12 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 82.39 |
| Shannon entropy | 0.36751 |
| G+C content | 0.31750 |
| Mean single sequence MFE | -24.79 |
| Consensus MFE | -12.36 |
| Energy contribution | -12.66 |
| Covariance contribution | 0.30 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.27 |
| SVM RNA-class probability | 0.987271 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 2721872 102 - 23011544 -AAUACAAAAUAAUAGAAGCACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCCAACUAUUGUGUGUCAUCCUUUUUUGUCCA--------- -...((((((.....((.((((((((..(((((...((((((((((((........)))))))))))))).-)))..)).)))))).))......))))))...--------- ( -28.20, z-score = -4.40, R) >droSim1.chr2L 2681259 102 - 22036055 -AAUACAAAAUAAUAGAAGCACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCCAACUAUUGUGUGUCAUCCUUUUUUGUCCA--------- -...((((((.....((.((((((((..(((((...((((((((((((........)))))))))))))).-)))..)).)))))).))......))))))...--------- ( -28.20, z-score = -4.40, R) >droSec1.super_5 888198 102 - 5866729 -AAUACAAAAUAAUAGAAGCACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCCAACUAUUGUGUGUCAUCCUUUUUUGUCCA--------- -...((((((.....((.((((((((..(((((...((((((((((((........)))))))))))))).-)))..)).)))))).))......))))))...--------- ( -28.20, z-score = -4.40, R) >droYak2.chr2L 2715525 104 - 22324452 -AAUACAAAAUAAUAGAAGCACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCCAACUAUUGUGUGUCAUCCUUUUUUGUGCACA------- -.................(((((((((((((.....((((((((((((........))))))))))))((.-(...((....))..).)))))))))))))))...------- ( -30.30, z-score = -4.38, R) >droEre2.scaffold_4929 2774767 102 - 26641161 -AAUACAAAAUAAUAGAAGCACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCCAACUAUUGUGUGUCAUCCUUUUUUGUCGA--------- -...((((((.....((.((((((((..(((((...((((((((((((........)))))))))))))).-)))..)).)))))).))......))))))...--------- ( -28.20, z-score = -4.24, R) >droWil1.scaffold_180772 1459486 103 - 8906247 -AGCACAAAAUAAUGUGUACA-AAAGAAGGACAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCAAACUAUUGUAUGUCAUCCUUUUUUCCUGUCG------- -.....((((..(((((((((-(.((..(((((...((((((((((((........)))))))))))))))-))...)).))))))).)))..)))).........------- ( -25.20, z-score = -3.63, R) >droAna3.scaffold_12984 183361 110 - 754457 -AGUACAAAAUAAUAGGAGAACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAGAAACAAUGU-CCCAACUAUUGUAUGUCAUCCUUUUUUAUGUUUUUAUACA- -.(((.(((((((.((((((((((((..(((((...((((((((((((........)))))))))))))).-)))..)).))))...)).))))...))))...))).))).- ( -25.50, z-score = -2.67, R) >droPer1.super_10 1056399 111 - 3432795 -AAUACGAAAUAAUAGGAGAACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-UCCAACUAUUGUGUGUCAUCGUUUUUUUUCCCAGGCUCCCA -.....((.(((((((..(((((..(.....)....((((((((((((........)))))))))))))))-))...)))))))...))........................ ( -21.80, z-score = -0.73, R) >dp4.chr4_group4 2050957 111 - 6586962 GGAUA-GAAAUAAUAGGAGAACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU-CCCAACUAUUGUGUGUCAUCGUUUUUUUUCCCAGGCUCCCA (((.(-((((..((.((...((((((..(((((...((((((((((((........)))))))))))))).-)))..)).))))...))))..)))))..))).......... ( -26.50, z-score = -1.81, R) >droVir3.scaffold_12963 11336286 95 - 20206255 -GACACAAAGUAACAC---AACAAAGAAGACCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUAG-CCCAACUAU----UGUCAUACUUUUCUUUUUG--------- -..............(---((.((((((((........((((((((((........))))))))))(((((-.....))))----).......)))))))))))--------- ( -16.50, z-score = -1.96, R) >droMoj3.scaffold_6500 20083452 95 - 32352404 -AACACAAAGUAUUGC---AACAAUGAAGGACAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUAG-CCCAACUAU----UGUCAUACUUUACUUCUCC--------- -.....(((((((.((---((.......((......((((((((((((........))))))))))))...-))......)----))).)))))))........--------- ( -20.52, z-score = -2.86, R) >droGri2.scaffold_15252 5455877 96 - 17193109 -GACACAAAAUAACAC---AACAAAGAAGAGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGGCCCCAACUAU----UGUCAUACUUUUCAUUUUG--------- -...............---......((((((......(((((((((((........)))))))))))(((((.........----.))))).))))))......--------- ( -18.30, z-score = -2.19, R) >consensus _AAUACAAAAUAAUAGAAGAACAAAGAAGGGCAAAUUUGUUUUUGAUAGAAAGUCAUGUCAAAAACAAUGU_CCCAACUAUUGUGUGUCAUCCUUUUUUGUCCA_________ ......................((((...((((...((((((((((((........)))))))))))).................))))...))))................. (-12.36 = -12.66 + 0.30)

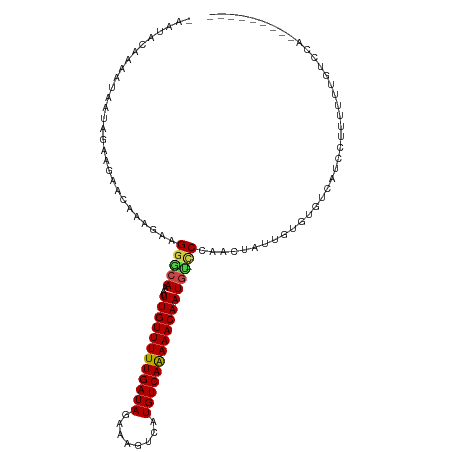
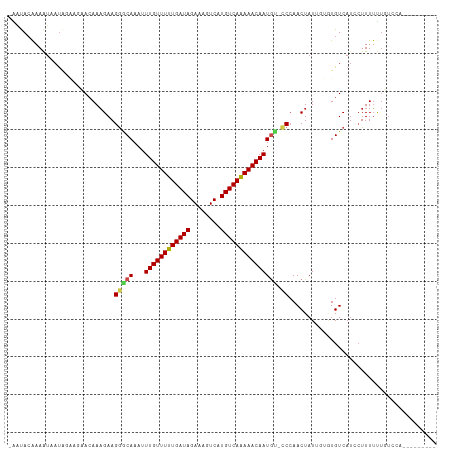
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:12:06 2011