
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 8,656,569 – 8,656,659 |
| Length | 90 |
| Max. P | 0.830614 |

| Location | 8,656,569 – 8,656,659 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 67.28 |
| Shannon entropy | 0.60416 |
| G+C content | 0.51272 |
| Mean single sequence MFE | -23.71 |
| Consensus MFE | -13.11 |
| Energy contribution | -12.49 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.59 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830614 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 8656569 90 - 21146708 --------ACCCCUAGCCAGGCGAA-UAGGAGCAGCUGCUCCCCGGAGCAUUGCUUUUGACUUGC---UUGAAUCCGUAC--------AAAAGUCCCCCUCUUUUUUUUU --------.........(((((((.-(((((((((.(((((....))))))))))))))..))))---))).........--------(((((.......)))))..... ( -25.40, z-score = -2.17, R) >droPer1.super_4 4249844 97 + 7162766 --------ACCCCUAAGCGAGAAAA--GGGAGCAGCUGCUGCCCGUGGCAUUGCUUUGGUCCUGC---UUGAAUCCGUACAACAGUCUGUGGGGACUGUCGCCCCUCCUU --------......(((.(((....--((((((....))).)))((((((..((.........))---.....(((.((((......)))).))).))))))..)))))) ( -26.60, z-score = 1.27, R) >dp4.chr3 14702347 97 - 19779522 --------ACCCCUAAGCGAGAAAA--GGGAGCAGCUGCUGCCCGUGGCAUUGCUUUGGUCCUGC---UUGAAUCCGUACAACAGUCUGUGGGGACUGUCGCCCCUCCUU --------......(((.(((....--((((((....))).)))((((((..((.........))---.....(((.((((......)))).))).))))))..)))))) ( -26.60, z-score = 1.27, R) >droAna3.scaffold_13266 6134655 100 - 19884421 ACCCCUAAAACUCUGUCAGGAUG----AGGAGCAGCUGCUCCCGGGAGCAUUGCUUUGGACUUGC---UUGAAUCCGUAC---ACACAAAAAGGACCCCUCCAGUUAUAU ........((((...(((((...----.(((((((.(((((....)))))))))))).......)---))))........---.........(((....))))))).... ( -23.20, z-score = 0.42, R) >droEre2.scaffold_4845 5437029 86 + 22589142 --------ACCCCUAGCCAAGCG----AGGAGCAGCUGCUCCCCGGAGCAUUGCUUUUGACUUGC---UUGAAUCCGUAC--------AAGAGUCCCCUCAUUUUUUUG- --------.........((((((----((((((((.(((((....)))))))))))....)))))---))).........--------..(((....))).........- ( -26.80, z-score = -3.07, R) >droYak2.chr2R 12845950 86 + 21139217 --------ACCCCUAGCCGAACG----AGGAGCAGCUGCUCCCCGGAGCAUUGCUUUUGACUUGC---UUGAAUCCGUAC--------AAAAGUCCCCCUUUUUUUUGG- --------........(((((.(----((((((((.(((((....))))))))))...(((((..---(((........)--------)))))))..))))...)))))- ( -22.70, z-score = -1.80, R) >droSec1.super_1 6194725 90 - 14215200 --------ACCCCUAGCCAGGCGAA-UAGGAGCAGCUGCUCCCCGGAGCAUUGCUUUUGACUUGC---UUGAAUCCGUAC--------AAAAGUCCCCCUUUUUUUUUUG --------.........(((((((.-(((((((((.(((((....))))))))))))))..))))---)))........(--------(((((...........)))))) ( -25.50, z-score = -2.44, R) >droSim1.chr2R 7136606 90 - 19596830 --------ACCCCUAGCCAGGCGAA-UAGGAGCAGCUGCUCCCCGGAGCAUUGCUUUUGACUUGC---UUGAAUCCGUAC--------AAAAGUCCCCCUUUUUUUGUUG --------.........(((((((.-(((((((((.(((((....))))))))))))))..))))---))).......((--------(((((........))))))).. ( -28.00, z-score = -3.04, R) >droWil1.scaffold_180708 12332846 92 - 12563649 --------UCCCUUAACAGUGCAACUCCUGAGCAGCUGUCGUUGGCAUUA--AACCAAAACUAACAAGUUGAAUCCCUAC--------AAAAGUUCCCCCUUUUUUUAAA --------.......(((((((.........))).)))).((.((.(((.--(((............))).))).)).))--------(((((......)))))...... ( -8.60, z-score = 1.00, R) >consensus ________ACCCCUAGCCGAGCGAA__AGGAGCAGCUGCUCCCCGGAGCAUUGCUUUUGACUUGC___UUGAAUCCGUAC________AAAAGUCCCCCUUUUUUUUUUU ............................(((((((.(((((....)))))))))))).(((((...........................)))))............... (-13.11 = -12.49 + -0.62)


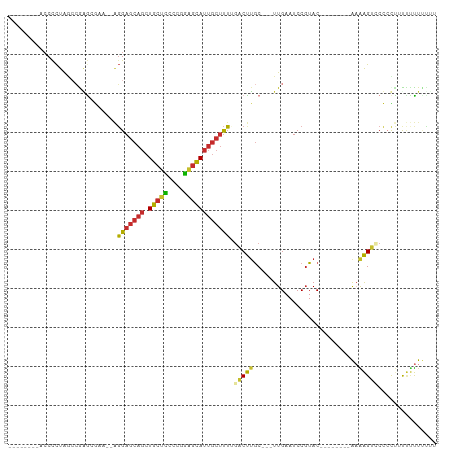
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:18:39 2011