
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 8,622,138 – 8,622,240 |
| Length | 102 |
| Max. P | 0.542356 |

| Location | 8,622,138 – 8,622,240 |
|---|---|
| Length | 102 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 77.13 |
| Shannon entropy | 0.41831 |
| G+C content | 0.37980 |
| Mean single sequence MFE | -19.91 |
| Consensus MFE | -8.66 |
| Energy contribution | -9.25 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.542356 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 8622138 102 - 21146708 -----UGCUUAUCGGAAAACUCCGAA-----AAACGAAACAAGAGG----CAAAGUUAUUGCACAAUGCGCAGAUAUAAGCCACACCAAGUAUAGUAAAUAUAGGCUAAGCGAAAC -----.((((((((((....))))).-----.............((----(.......((((.......))))......)))........................)))))..... ( -17.32, z-score = -0.66, R) >droSim1.chr2R 7103050 89 - 19596830 -----CGCUUAUCGGAAAACUCCGAA-----AAACGAAACAAGAGG----CAAAAUUAUUGCACAAUACAC-------------ACCAAGUAUAGUAAAUAUAGGCUAAGCGAAAC -----(((((.(((((....))))).-----.............((----(.....((((..((.((((..-------------.....)))).)).))))...)))))))).... ( -18.00, z-score = -3.50, R) >droSec1.super_1 6160288 102 - 14215200 -----CGCUUAUCGGAAAACUCCGAA-----AAACGAAACAAGAGG----CAAAUUUAUUGCACAAUGCGCAGAUAUAAACCAAACCAAGUAUAGUAAAUAUAGGCUAAGCGAAAC -----(((((.(((((....))))).-----.............((----(.....((((..((.((((....................)))).)).))))...)))))))).... ( -17.35, z-score = -1.51, R) >droYak2.chr2R 12810818 100 + 21139217 -----CGCUUAUCGGAAAACUCCGAA-----AAGCGAAACAAGAGG----CAAAGUUAUUGCACAAUGCA--AAUAUAUGCCGCACCAAGUAUAGUAAAUAUAGGCUAAGCAAAAC -----(((((.(((((....))))).-----)))))......(.((----((......((((.....)))--).....)))).).....((.((((........)))).))..... ( -26.10, z-score = -3.77, R) >droEre2.scaffold_4845 5402543 105 + 22589142 -----CGCUUAUCGGAAAACGCCGAAAAACGAAACGAAACAAGAGG----CAAAGUUAUUGCACAAUGCA--AAUAUAUACCGCACCAAGUAUAGUAAAUAUAGGCUAAGCGAAAC -----(((((.((((......))))...................((----(.......((((.....)))--)...(((((........)))))..........)))))))).... ( -18.50, z-score = -1.17, R) >droAna3.scaffold_13266 11365605 113 + 19884421 CUCCGGGCUUAUCGCCAGGCUUACAA--AAAAAACUAAAUAUCAAGUGACCAAAAUAAUUACACAAUGCAU-AAAAUAUAUGCUACAGGGUAUAGUAAGUAUAUGCCAAGGCACAC ..((.(((.....))).)).......--.................(((.((................((((-(.....))))).....((((((.......))))))..))))).. ( -22.20, z-score = -1.34, R) >consensus _____CGCUUAUCGGAAAACUCCGAA_____AAACGAAACAAGAGG____CAAAGUUAUUGCACAAUGCAC_AAUAUAUACCACACCAAGUAUAGUAAAUAUAGGCUAAGCGAAAC .....((((((((((......))))...............................((((..((.((((....................)))).)).)))).....)))))).... ( -8.66 = -9.25 + 0.59)

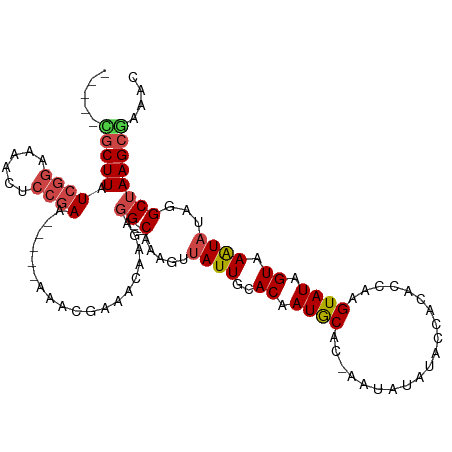

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:18:35 2011