
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 8,498,404 – 8,498,471 |
| Length | 67 |
| Max. P | 0.848316 |

| Location | 8,498,404 – 8,498,471 |
|---|---|
| Length | 67 |
| Sequences | 7 |
| Columns | 74 |
| Reading direction | forward |
| Mean pairwise identity | 79.16 |
| Shannon entropy | 0.39111 |
| G+C content | 0.53285 |
| Mean single sequence MFE | -13.83 |
| Consensus MFE | -6.94 |
| Energy contribution | -6.86 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.57 |
| SVM RNA-class probability | 0.745599 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 8498404 67 + 21146708 --AUACACAUUCCUUUGAGCUU---UGCACGCGCUAGUU-UCCCCCUCCGCGCCA-AAUGCAGAUCUCAGCACU --............(((((.((---((((.((((.....-.........))))..-..)))))).))))).... ( -15.54, z-score = -2.73, R) >droAna3.scaffold_13266 13096810 70 + 19884421 AUAUAUAUCCUUCCUUGAGCUU---UGCACGCGCUAGUC-UCCCCCACCGCGCCACAAUGCAGACCUCUGCACU ................(((.((---((((.((((.....-.........)))).....)))))).)))...... ( -13.14, z-score = -1.80, R) >droEre2.scaffold_4845 5288287 67 - 22589142 --GUACACAUAUCCUCGAGCUU---UGCACGCGCUAGUU-CCCCCCUCCGCGCCA-AAUGCAGAUCUCCGCACU --..............(((.((---((((.((((.....-.........))))..-..)))))).)))...... ( -13.74, z-score = -1.90, R) >droYak2.chr2R 12693456 67 - 21139217 --AUACACAUAUCCUCGAGCUU---UGCACGCGCUAGUU-UCCCCCUCCGCGCCA-AAUGCAGAUCUCCGCACU --..............(((.((---((((.((((.....-.........))))..-..)))))).)))...... ( -13.74, z-score = -2.07, R) >droSec1.super_1 6044473 67 + 14215200 --AUACACACUCCCUCGAGCUU---UGCACGCGCUAGUU-UCCCCCUCCGCGCCA-AAUGCAGAUCUCCGCACU --..............(((.((---((((.((((.....-.........))))..-..)))))).)))...... ( -13.74, z-score = -2.17, R) >droSim1.chr2R 6982958 67 + 19596830 --AUACACACUCCCUCGAGCUU---UGCACGCGCUAGUU-UCCCCCUCCGCGCCA-AAUGCAGAUCUCCGCACU --..............(((.((---((((.((((.....-.........))))..-..)))))).)))...... ( -13.74, z-score = -2.17, R) >droGri2.scaffold_15245 4234392 71 + 18325388 --AUAUUUAUCCCAUUUAUGUUAUACGCAUGCGCUAGUUGCCCCUCUAAGUGCUA-UGUGAUAAGAAUUUCACU --................(.((((.((((((((((((........)).))))).)-)))))))).)........ ( -13.20, z-score = -1.80, R) >consensus __AUACACAUUCCCUCGAGCUU___UGCACGCGCUAGUU_UCCCCCUCCGCGCCA_AAUGCAGAUCUCCGCACU ..................((......))..((((...............)))).....(((........))).. ( -6.94 = -6.86 + -0.08)


| Location | 8,498,404 – 8,498,471 |
|---|---|
| Length | 67 |
| Sequences | 7 |
| Columns | 74 |
| Reading direction | reverse |
| Mean pairwise identity | 79.16 |
| Shannon entropy | 0.39111 |
| G+C content | 0.53285 |
| Mean single sequence MFE | -19.93 |
| Consensus MFE | -14.96 |
| Energy contribution | -14.74 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.848316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
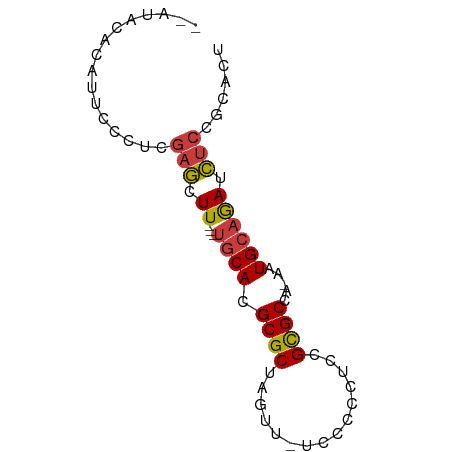
>dm3.chr2R 8498404 67 - 21146708 AGUGCUGAGAUCUGCAUU-UGGCGCGGAGGGGGA-AACUAGCGCGUGCA---AAGCUCAAAGGAAUGUGUAU-- .....((((...(((((.-..((((.....((..-..)).)))))))))---...)))).............-- ( -19.30, z-score = -0.80, R) >droAna3.scaffold_13266 13096810 70 - 19884421 AGUGCAGAGGUCUGCAUUGUGGCGCGGUGGGGGA-GACUAGCGCGUGCA---AAGCUCAAGGAAGGAUAUAUAU (((((((....)))))))((.(((((((..((..-..)).)).))))).---..)).................. ( -22.60, z-score = -2.15, R) >droEre2.scaffold_4845 5288287 67 + 22589142 AGUGCGGAGAUCUGCAUU-UGGCGCGGAGGGGGG-AACUAGCGCGUGCA---AAGCUCGAGGAUAUGUGUAC-- .(((((.(.((((((.((-(((((((....((..-..))..))))).))---))))....)))).).)))))-- ( -19.30, z-score = -0.72, R) >droYak2.chr2R 12693456 67 + 21139217 AGUGCGGAGAUCUGCAUU-UGGCGCGGAGGGGGA-AACUAGCGCGUGCA---AAGCUCGAGGAUAUGUGUAU-- .(((((.(.((((((.((-(((((((....((..-..))..))))).))---))))....)))).).)))))-- ( -19.60, z-score = -0.83, R) >droSec1.super_1 6044473 67 - 14215200 AGUGCGGAGAUCUGCAUU-UGGCGCGGAGGGGGA-AACUAGCGCGUGCA---AAGCUCGAGGGAGUGUGUAU-- (((((((....)))))))-..((((.....((..-..)).))))(((((---..((((....)))).)))))-- ( -24.00, z-score = -2.45, R) >droSim1.chr2R 6982958 67 - 19596830 AGUGCGGAGAUCUGCAUU-UGGCGCGGAGGGGGA-AACUAGCGCGUGCA---AAGCUCGAGGGAGUGUGUAU-- (((((((....)))))))-..((((.....((..-..)).))))(((((---..((((....)))).)))))-- ( -24.00, z-score = -2.45, R) >droGri2.scaffold_15245 4234392 71 - 18325388 AGUGAAAUUCUUAUCACA-UAGCACUUAGAGGGGCAACUAGCGCAUGCGUAUAACAUAAAUGGGAUAAAUAU-- .((((........)))).-..(((....(..((....))..)...))).(((..((....))..))).....-- ( -10.70, z-score = -0.25, R) >consensus AGUGCGGAGAUCUGCAUU_UGGCGCGGAGGGGGA_AACUAGCGCGUGCA___AAGCUCGAGGGAAUGUGUAU__ .((((((....))))))....((((((..((......))..).))))).......................... (-14.96 = -14.74 + -0.22)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:18:22 2011