| Sequence ID | dm3.chr2R |
|---|---|
| Location | 8,287,373 – 8,287,502 |
| Length | 129 |
| Max. P | 0.953873 |

| Location | 8,287,373 – 8,287,502 |
|---|---|
| Length | 129 |
| Sequences | 7 |
| Columns | 139 |
| Reading direction | forward |
| Mean pairwise identity | 66.98 |
| Shannon entropy | 0.65330 |
| G+C content | 0.31450 |
| Mean single sequence MFE | -26.18 |
| Consensus MFE | -13.02 |
| Energy contribution | -12.72 |
| Covariance contribution | -0.29 |
| Combinations/Pair | 1.69 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.953873 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
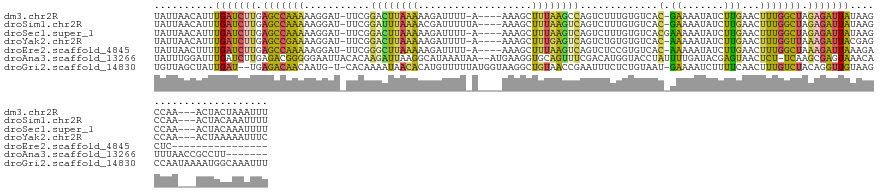
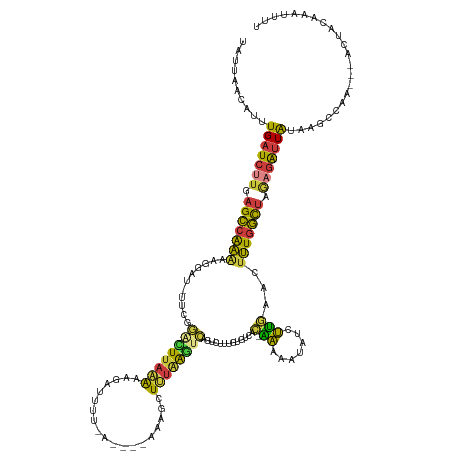
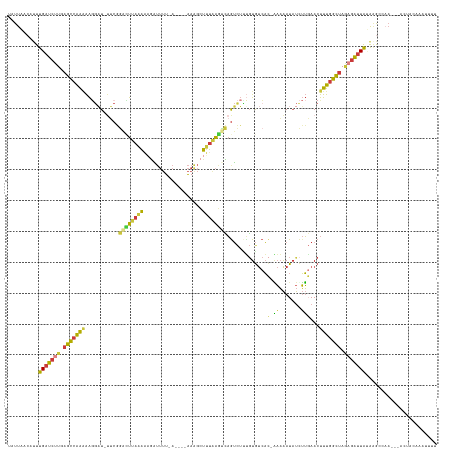
>dm3.chr2R 8287373 129 + 21146708 UAUUAACAUUUGAUCUUGAGCCAAAAAGGAU-UUCGGACUUAAAAAGAUUUU-A----AAAGCUUUAAGCCAGUCUUUGUGUCAC-GAAAAUAUCUUGAACUUUGGCUAGAGAUUAUAAGCCAA---ACUACUAAAUUU ..........(((((((.(((((((.(((((-(((((((....(((((((..-.----...((.....)).)))))))..))).)-)))...)))))....))))))).)))))))........---............ ( -29.20, z-score = -2.61, R) >droSim1.chr2R 6758654 130 + 19596830 UAUUAACAUUUGAUCUUGAGCCAAAAAGGAU-UUCGGAUUUAAAACGAUUUUUA----AAAGCUUUAAGUCAGUCUUUGUGUCAC-GAAAAUAUCUUGAACUUUGGCUAGAGAUUAUAAGCCAA---ACUACAAAUUUU ..........(((((((.(((((((.(((((-(((((((.(((((((((((...----........))))).)).)))).))).)-)))...)))))....))))))).)))))))........---............ ( -26.40, z-score = -1.77, R) >droSec1.super_1 5832189 130 + 14215200 UAUUAACAUUUGAUCUUGAGCCAAAAAGGAU-UUCGGACUUAAAAAGAUUUU-A----AAAGCUUUAAGUCAGUCUUUGUGUCACGAAAAAUAUCUUGAACUUUGGCUAGAGAUUAUAAGCCAA---ACUACAAAUUUU ..........(((((((.(((((((.(((((-(((((((....(((((((((-(----(.....))))...)))))))..))).))))....)))))....))))))).)))))))........---............ ( -28.30, z-score = -2.29, R) >droYak2.chr2R 12480511 129 - 21139217 UAUUAACAUUUGAUCUUGAGCCGAAAAGGAU-UUCGGACUUAAAAAGAUUUU-A----AAAGCUUUGAGUCAGUCUGUGUGUCAC-AAAAAUAUCUUGAACUUUGGUUAAAGAUUACGAGCCAA---ACUAAAAAUUUC ..........(((((((.(((((((.(((((-....((((((((........-.----.....))))))))....((((...)))-).....)))))....))))))).)))))))........---............ ( -22.34, z-score = -0.05, R) >droEre2.scaffold_4845 5083740 116 - 22589142 UAUUAACUUUUGAUCUUGAGCCAAAAAGGAU-UUCGGGCUUAAAAAGAUUUU-A----AAAGCUUUAAGUCAGUCUCCGUGUCAC-AAAAAUAUCUUGAACUUUGGCUAAAGAUUAAAGACUC---------------- .......((((((((((.(((((((..(((.-....((((((((........-.----.....))))))))....)))......(-((.......)))...))))))).))))))))))....---------------- ( -26.44, z-score = -1.98, R) >droAna3.scaffold_13266 5582547 129 + 19884421 UAUUUGGAUUUGAUCUUGAGACGGGGGAAUUACACAAGAUUAAGGCAUAAAUAA--AUGAAGGUGCAGUUUCGACAUGGUACCUAUUUUGAUACGAGUAACUCU-UCAAGCGAGUAAACAUUUAACCGCCUU------- ((((((..((((((((((................))))))))))...)))))).--.(((((((((.((.((((.((((...)))).)))).))..)).)).))-))).(((..(((....)))..)))...------- ( -24.69, z-score = -0.08, R) >droGri2.scaffold_14830 698779 134 - 6267026 UGUUAGCUAUUGAU--UGAGACAACAAUG-U-CACAAAAUAACACAUGUUUUUAUGGUAAGGCUGUAACCGAAUUUCUCUGUAAU-GAAAAUCUUUUCAACUUUGUCUACAGGUUGUAAGCCAAUAAAAUGGCAAAUUU .....((((((..(--((.((((....))-)-).)))....((.((((....))))))..((((..((((((.((((........-)))).))..........((....))))))...)))).....))))))...... ( -25.90, z-score = -0.55, R) >consensus UAUUAACAUUUGAUCUUGAGCCAAAAAGGAU_UUCGGACUUAAAAAGAUUUU_A____AAAGCUUUAAGUCAGUCUUUGUGUCAC_AAAAAUAUCUUGAACUUUGGCUAGAGAUUAUAAGCCAA___ACUACAAAUUUU ..........(((((((.(((((((...........((((((((...................)))))))).......((((........)))).......))))))).)))))))....................... (-13.02 = -12.72 + -0.29)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:17:51 2011