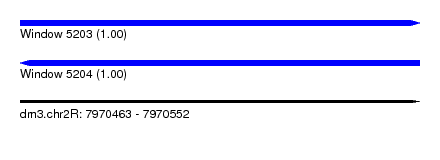

| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,970,463 – 7,970,552 |
| Length | 89 |
| Max. P | 0.999315 |

| Location | 7,970,463 – 7,970,552 |
|---|---|
| Length | 89 |
| Sequences | 4 |
| Columns | 89 |
| Reading direction | forward |
| Mean pairwise identity | 66.67 |
| Shannon entropy | 0.53792 |
| G+C content | 0.53151 |
| Mean single sequence MFE | -32.18 |
| Consensus MFE | -16.33 |
| Energy contribution | -18.82 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.31 |
| Mean z-score | -3.20 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.79 |
| SVM RNA-class probability | 0.999315 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
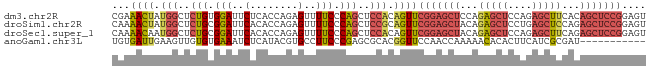
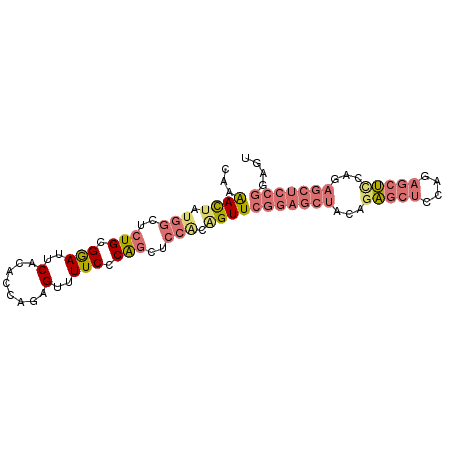
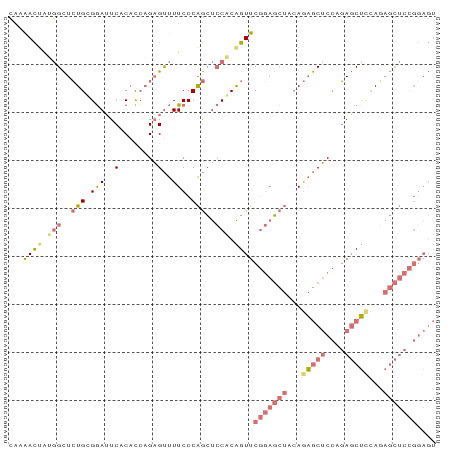
>dm3.chr2R 7970463 89 + 21146708 CGAAACUAUGGCUCUGUGGAUUCUCACCAGAGUUUUCCCAGCUCCACAGUUCGGAGCUCCAGAGCUCCAGAGCUUCACAGCUCCGGAGU ...((((.(((..(((.(((..(((....)))...))))))..))).))))(((((((...(((((....)))))...))))))).... ( -34.10, z-score = -2.82, R) >droSim1.chr2R 6507481 89 + 19596830 CAAAACUAUGGCUCUGCGGAUUCACACCAGAGUUUUCCCAGCUCCGCAGUUCGGAGCUACAGAGCUCCUGAGCUCCAGAGCUCCGGAGU .........(((((((.(........))))))))......((((((.(((((((((((.(((.....))))))))).))))).)))))) ( -37.40, z-score = -3.56, R) >droSec1.super_1 5519477 89 + 14215200 CAAAACAAUGGCUCUGCGGAUUCACACCAGAGUUUUCCCAGCUCCACAGUUCGGAGCUACAGAGCUCCAGAGCUUCAGAGCUCCGGAGU .........(((((((.(........))))))))((((.(((((...(((((((((((....)))))).)))))...)))))..)))). ( -32.90, z-score = -3.25, R) >anoGam1.chr3L 8329706 78 - 41284009 UGUGAUUGAAGUUGUGUGAAAUCUCAUACGUGCCUUCCCGAGCGCACGGUUCCAACCAAAAACACACUUCAUCGCGAU----------- .(((((.(((((.((((............(((((((...))).))))(((....)))....))))))))))))))...----------- ( -24.30, z-score = -3.19, R) >consensus CAAAACUAUGGCUCUGCGGAUUCACACCAGAGUUUUCCCAGCUCCACAGUUCGGAGCUACAGAGCUCCAGAGCUCCAGAGCUCCGGAGU ...((((.(((..(((.(((..(........)..))).)))..))).))))(((((((...(((((....)))))...))))))).... (-16.33 = -18.82 + 2.50)
| Location | 7,970,463 – 7,970,552 |
|---|---|
| Length | 89 |
| Sequences | 4 |
| Columns | 89 |
| Reading direction | reverse |
| Mean pairwise identity | 66.67 |
| Shannon entropy | 0.53792 |
| G+C content | 0.53151 |
| Mean single sequence MFE | -35.00 |
| Consensus MFE | -22.05 |
| Energy contribution | -25.18 |
| Covariance contribution | 3.13 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.66 |
| SVM RNA-class probability | 0.999128 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
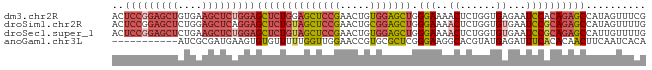
>dm3.chr2R 7970463 89 - 21146708 ACUCCGGAGCUGUGAAGCUCUGGAGCUCUGGAGCUCCGAACUGUGGAGCUGGGAAAACUCUGGUGAGAAUCCACAGAGCCAUAGUUUCG .((((((((((....))))))))))....((....))(((((((((..((((((...(((....)))..))).)))..))))))))).. ( -42.30, z-score = -4.20, R) >droSim1.chr2R 6507481 89 - 19596830 ACUCCGGAGCUCUGGAGCUCAGGAGCUCUGUAGCUCCGAACUGCGGAGCUGGGAAAACUCUGGUGUGAAUCCGCAGAGCCAUAGUUUUG ....(((((((..((((((....))))))..)))))))..(((((((((..((.....))..)).....)))))))............. ( -38.30, z-score = -2.56, R) >droSec1.super_1 5519477 89 - 14215200 ACUCCGGAGCUCUGAAGCUCUGGAGCUCUGUAGCUCCGAACUGUGGAGCUGGGAAAACUCUGGUGUGAAUCCGCAGAGCCAUUGUUUUG ..(((((((((....)))))))))((((((((((((((.....))))))).(((..((......))...)))))))))).......... ( -39.10, z-score = -3.92, R) >anoGam1.chr3L 8329706 78 + 41284009 -----------AUCGCGAUGAAGUGUGUUUUUGGUUGGAACCGUGCGCUCGGGAAGGCACGUAUGAGAUUUCACACAACUUCAAUCACA -----------...(.((((((((((((....((((.....(((((.((.....))))))).....))))..)))).))))).))).). ( -20.30, z-score = -0.19, R) >consensus ACUCCGGAGCUCUGAAGCUCAGGAGCUCUGUAGCUCCGAACUGUGGAGCUGGGAAAACUCUGGUGAGAAUCCACAGAGCCAUAGUUUCG ..(((((((((....)))))))))((((((((((((((.....))))))).(((..((......))...)))))))))).......... (-22.05 = -25.18 + 3.13)
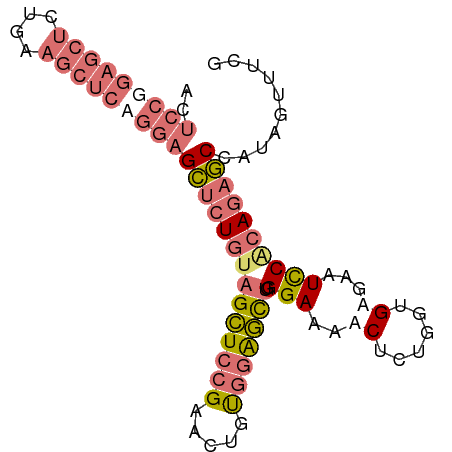
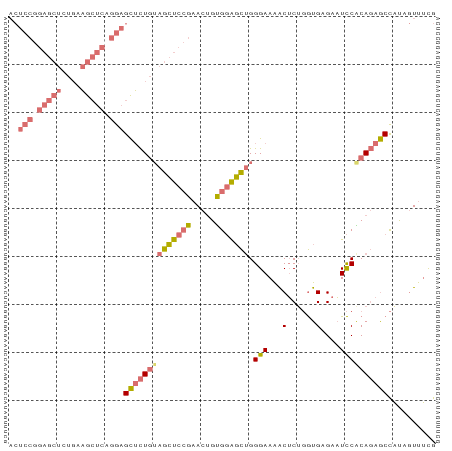
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:16:56 2011