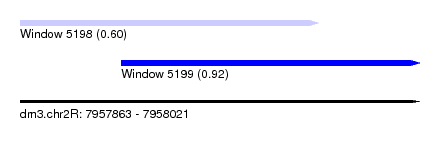
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,957,863 – 7,958,021 |
| Length | 158 |
| Max. P | 0.920089 |

| Location | 7,957,863 – 7,957,981 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 93.46 |
| Shannon entropy | 0.11204 |
| G+C content | 0.35282 |
| Mean single sequence MFE | -29.29 |
| Consensus MFE | -20.94 |
| Energy contribution | -21.74 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.600578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
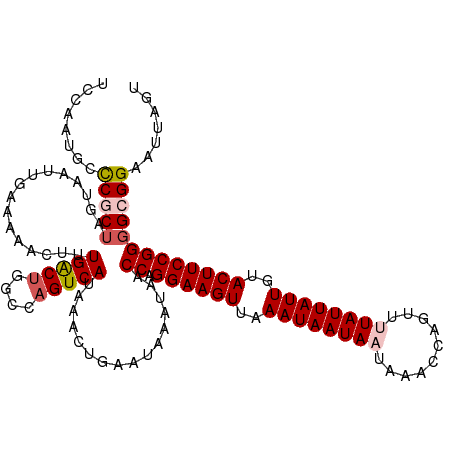
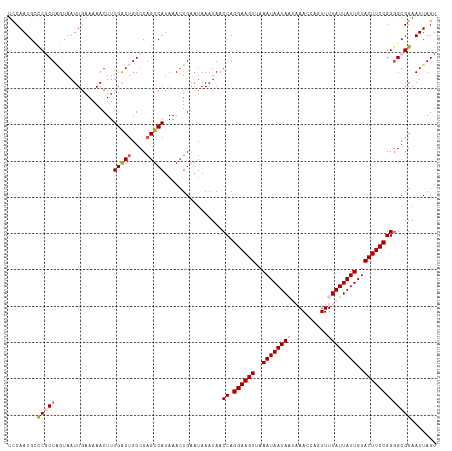
>dm3.chr2R 7957863 118 + 21146708 UCCAAUGGCCGCUGGUAAUUGAAAAACUUUGACUGGCCAGUCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGU ........((((((((.(((.........(((((....)))))...........))).))).((((((...(((((((((((.....)))))))))))))))))..)))))....... ( -30.55, z-score = -2.38, R) >droSim1.chr2R 6495063 118 + 19596830 UCCAAUGGCCGCUAGUAAUUGAAAAACUUUGACUGGCCAGUCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGU (((.(((((.((((((....(.....)....))))))..)))))...............((.((((((...(((((((((((.....))))))))))))))))).))..)))...... ( -30.80, z-score = -2.81, R) >droSec1.super_1 5507183 118 + 14215200 UUCAAUGCCCGCUAGUAAUUGAAAAACUUUGACUGGCCAGUCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGU .....(((((((((((....(.....)....))))))((((.....))))............((((((...(((((((((((.....))))))))))))))))).)))))........ ( -31.30, z-score = -3.34, R) >droYak2.chr2R 12144323 115 - 21139217 UCCAAUGCUCACUAGUAAUUGAAAAACUUUGACUGGCCAGUCAUAAACUGAAUAAAUAACCAGGAAGUGAAAUAAUA---AACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGU (((...((((..((((..((....))...(((((....)))))...))))............(((((((.(((((((---((.....))))))))).))))))).)))))))...... ( -26.80, z-score = -2.00, R) >droEre2.scaffold_4845 4749604 115 - 22589142 GCCAAUGCUCACUUGUAAUUGAAAAACUUUGGCUGGCCCGUCAUAAAGUGCAUAAAUAACCAGGAAGUGAAAUAAUA---AACCAGUUUUAUUAUUGUACUUCCGGUGCGGAAUUAGU .((.(((((((........)))...((((((((......))))..)))))))).....(((.(((((((.(((((((---((.....))))))))).))))))))))..))....... ( -27.00, z-score = -1.36, R) >consensus UCCAAUGCCCGCUAGUAAUUGAAAAACUUUGACUGGCCAGUCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGU ........(((((................(((((....)))))................((.((((((..((((((((..........))))))))..)))))))))))))....... (-20.94 = -21.74 + 0.80)

| Location | 7,957,903 – 7,958,021 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 95.41 |
| Shannon entropy | 0.07785 |
| G+C content | 0.30153 |
| Mean single sequence MFE | -25.56 |
| Consensus MFE | -18.92 |
| Energy contribution | -19.52 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.99 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.920089 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
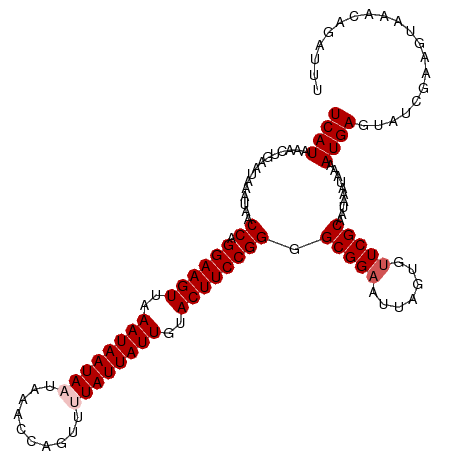
>dm3.chr2R 7957903 118 + 21146708 UCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGUGUUCGCAAUAAAUAAAAUGAGUAUCGAAGUAAACAGAUUU .......(((.........((.((((((...(((((((((((.....))))))))))))))))).))(((((.......)))))...........................))).... ( -24.30, z-score = -2.69, R) >droSim1.chr2R 6495103 118 + 19596830 UCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGUGCUCGCAAUAAAUAAAAUGAGUAUCGAAGUAAACAGAUUU .......(((.........((.((((((...(((((((((((.....))))))))))))))))).))......(..(((((((............)))))))..)......))).... ( -24.90, z-score = -2.80, R) >droSec1.super_1 5507223 118 + 14215200 UCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGUGUUCGCAAUAAAUAAAAUGAGUAUCGAAGUAAACAGAUUU .......(((.........((.((((((...(((((((((((.....))))))))))))))))).))(((((.......)))))...........................))).... ( -24.30, z-score = -2.69, R) >droYak2.chr2R 12144363 115 - 21139217 UCAUAAACUGAAUAAAUAACCAGGAAGUGAAAUAAUA---AACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGUGUUCGCAAUAAAUAGAAUGAGUAUCGAAGUAAACAGAUUU .......(((.........((.(((((((.(((((((---((.....))))))))).))))))).))(((((.......)))))...........................))).... ( -25.40, z-score = -2.73, R) >droEre2.scaffold_4845 4749644 115 - 22589142 UCAUAAAGUGCAUAAAUAACCAGGAAGUGAAAUAAUA---AACCAGUUUUAUUAUUGUACUUCCGGUGCGGAAUUAGUGUUCGCAAUAAAUAGCAUGAGUAUCGAAGUAAACAGAUAU .......((((.......(((.(((((((.(((((((---((.....))))))))).))))))))))(((((.......)))))........))))..(((((..........))))) ( -28.90, z-score = -4.05, R) >consensus UCAUAAACUGAAUAAAUAACCAGGAAGUUAAAUAAUAAUAAACCAGUUUUAUUAUUGUACUUCCGGGGCGGAAUUAGUGUUCGCAAUAAAUAAAAUGAGUAUCGAAGUAAACAGAUUU ((((...............((.((((((..((((((((..........))))))))..)))))))).(((((.......)))))..........)))).................... (-18.92 = -19.52 + 0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:16:52 2011