
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,951,746 – 7,951,851 |
| Length | 105 |
| Max. P | 0.712092 |

| Location | 7,951,746 – 7,951,851 |
|---|---|
| Length | 105 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 68.88 |
| Shannon entropy | 0.62376 |
| G+C content | 0.45194 |
| Mean single sequence MFE | -22.88 |
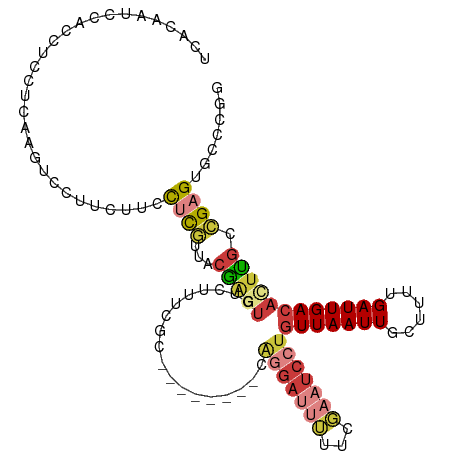
| Consensus MFE | -9.24 |
| Energy contribution | -10.66 |
| Covariance contribution | 1.41 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.712092 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
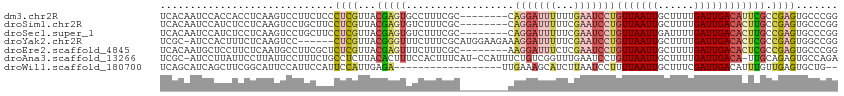
>dm3.chr2R 7951746 105 - 21146708 UCACAAUCCACCACCUCAAGUCCUUCUCCCUCGUUACGAGUGCCUUUCGC--------CAGGAUUUUUUGAAUCCUGUUAAUUGCUUUUGAUUGACAUUCGCCGAGUGCCCGG .............................((((...((((((........--------((((((((...))))))))((((((......)))))))))))).))))....... ( -19.30, z-score = -0.66, R) >droSim1.chr2R 6488724 105 - 19596830 UCACAAUCCAUCUCCUCAAGUCCUGCUUCCUCGUUACGAGUGUCUUUCGC--------CAGGAUUUUUCGAAUCCUGUUAAUUGCUUUUGAUUGACACUUGCCGAGUGCCCGG .....................((.((...((((...((((((((..((((--------((((((((...))))))))......))....))..)))))))).)))).))..)) ( -25.20, z-score = -2.16, R) >droSec1.super_1 5501054 105 - 14215200 UCACAAUCCAUCUCCUCAAGUCCUGCUUCCUCGUUACGAGUGUCUUUCGC--------CAGGAUUUUUCGAAUCCUGUUAAUUGAUUUUGAUUGACACUUGCCGAGUGCCCGG .....................((.((...((((...((((((((..(((.--------((((((((...))))))))...........)))..)))))))).)))).))..)) ( -24.80, z-score = -1.94, R) >droYak2.chr2R 12138377 106 + 21139217 UCGC-AUCCACUUUCUCAAGUCC------CUCGUUACGGGUUUCUUUCGCAUGGAAGAAAGGAUUUUUCGAAUCCUGUUAAUUGCUUUUGAUUGACACUCGCCGAGUGGCCGG ....-..............(((.------((((...((((((((((((....)))))))(((((((...)))))))(((((((......)))))))))))).)))).)))... ( -27.10, z-score = -0.50, R) >droEre2.scaffold_4845 4743877 105 + 22589142 UCACAAUGCUCCUUCUCAAUGCCUUCGCUCUCGUUACGAGUUUCUUUCGC--------AAGGAUUUCUCGAAUCCUGUUAAUUGCUUUUGAUUGACACUCGCCGAGUGCCCGG .....................((...((.((((...(((((.........--------.(((((((...)))))))(((((((......)))))))))))).)))).))..)) ( -23.70, z-score = -1.27, R) >droAna3.scaffold_13266 17209556 110 + 19884421 UCGC-AUCCUUAUUCCUUAUUCCUUUCUGCCUCUUACACUUUCCACUUUCAU-CCAUUUCUGUCGGUUUGAAUCCUGUUAAUUGCUUUUGAUUGACA-UUGCAGAGUGCCAGA ..((-((..................(((((.................((((.-((.........))..))))...((((((((......))))))))-..))))))))).... ( -15.16, z-score = -0.49, R) >droWil1.scaffold_180700 901358 93 + 6630534 UCAGCAUCAGCUUCGGCAUUCCAUUCCAUUCCAUUGAGA------------------UUGAAAGCAUCUUAAUCCUUUUAAUUGCUUUCGAUUGACAUUUGUUGAGUGCUG-- .((((((((((....)).................((.((------------------(((((((((..((((.....)))).)))))))))))..)).....))).)))))-- ( -24.90, z-score = -2.81, R) >consensus UCACAAUCCACCUCCUCAAGUCCUUCUUCCUCGUUACGAGUGUCUUUCGC________CAGGAUUUUUCGAAUCCUGUUAAUUGCUUUUGAUUGACACUUGCCGAGUGCCCGG .............................((((...(((((..................(((((((...)))))))(((((((......)))))))))))).))))....... ( -9.24 = -10.66 + 1.41)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:16:47 2011