| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,905,444 – 7,905,521 |
| Length | 77 |
| Max. P | 0.948444 |

| Location | 7,905,444 – 7,905,521 |
|---|---|
| Length | 77 |
| Sequences | 12 |
| Columns | 79 |
| Reading direction | forward |
| Mean pairwise identity | 76.54 |
| Shannon entropy | 0.52201 |
| G+C content | 0.44544 |
| Mean single sequence MFE | -18.28 |
| Consensus MFE | -10.81 |
| Energy contribution | -11.07 |
| Covariance contribution | 0.26 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.948444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
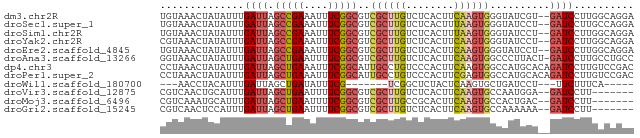
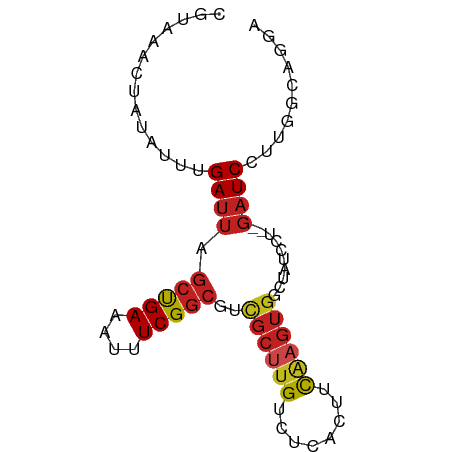
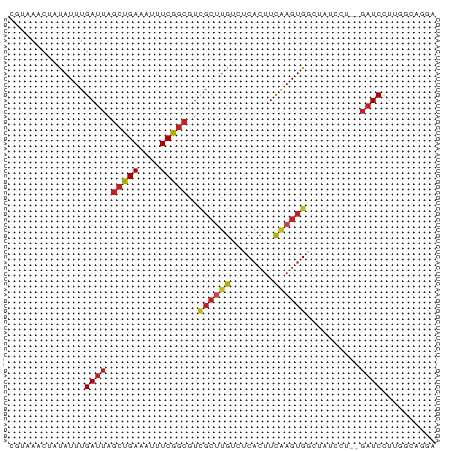
>dm3.chr2R 7905444 77 + 21146708 UGUAAACUAUAUUUGAUUAGCCGAAAUUUCGGCGUCGCUUGUCUCACUUCAAGUGGGUAUCGU--GAUCCUUGGCAGGA ...................(((((....)))))(((((....(((((.....)))))....))--)))(((....))). ( -19.20, z-score = -0.83, R) >droSec1.super_1 5455477 77 + 14215200 UGUAAACUAUAUUUGAUUAGCCGAAAUUUCGGCGUCGCUUGUCUCACUUUAAGUGGGUAUCCU--GAUCCUUGCCAGGA .....(((...........(((((....)))))..((((((........))))))))).((((--(........))))) ( -19.30, z-score = -1.37, R) >droSim1.chr2R 6441980 77 + 19596830 UGUAAACUAUAUUUGAUUAGCCGAAAUUUCGGCGUCGCUUGUCUCACUUUAAGUGGGUAUCCU--GAUCCUUGGCAGGA .....(((...........(((((....)))))..((((((........))))))))).((((--(........))))) ( -19.10, z-score = -1.01, R) >droYak2.chr2R 12091264 77 - 21139217 CGUAAACUAUAUUUGAUUAGCCGAAAUUUCGGCGUCGCUUGUCUCACUUCAAGUGGGUAUCCU--GAUCCUUGGCAGGA .....(((...........(((((....)))))..((((((........))))))))).((((--(........))))) ( -20.60, z-score = -1.35, R) >droEre2.scaffold_4845 4698129 77 - 22589142 UGUAAACUAUAUUUGAUUAGCCGAAAUUUCGGCGUCGCUUGUCUCACUUCAAGUGGGUAUCCU--GAUCCUUGGCAGGA .....(((...........(((((....)))))..((((((........))))))))).((((--(........))))) ( -20.60, z-score = -1.41, R) >droAna3.scaffold_13266 17162023 78 - 19884421 GGUAAACUAUAUUUGAUUAGCUGAAUUUUCGGCGUCGCUUGUCUCACUUCAAGUGGCCCUUACU-GAUCCUUGCCUGCC (((((..............(((((....)))))((((((((........)))))))).......-.....))))).... ( -20.60, z-score = -2.59, R) >dp4.chr3 13189647 79 - 19779522 CCUAAACUAUAUUUGAUUAGCUGAAAUUUCGGCAUUGCCUGUCCCACUUCAAGUGGCCAUGCACAGAUCCUUGUCCGAC ..........(((((....(((((....)))))..(((.((..((((.....)))).)).))))))))........... ( -16.30, z-score = -1.11, R) >droPer1.super_2 2225561 79 - 9036312 CCUAAACUAUAUUUGAUUAGCUGAAAUUUCGGCAUUGCCUGUCCCACUUCGAGUGGCCAUGCACAGAUCCUUGUCCGAC ..........(((((....(((((....)))))..(((.((..((((.....)))).)).))))))))........... ( -16.30, z-score = -0.94, R) >droWil1.scaffold_180700 844432 61 - 6630534 ---AACCUACAUUUGAUUAGCUGAUAUUUCG-------UCGGCUCUACUCAAGUGCUGAUCCU---UUCUUUCA----- ---...(..(((((((..(((((((.....)-------))))))....)))))))..).....---........----- ( -13.70, z-score = -3.15, R) >droVir3.scaffold_12875 12010020 70 - 20611582 CGUCAACUGCAUUUGAUUAGCUGAAUUUUCGGCGUCGCUUGUCUCACUUCAAGUGCCAAUGGA--GAUCCUU------- ......((.((((......(((((....)))))(.((((((........)))))).))))).)--)......------- ( -16.50, z-score = -0.71, R) >droMoj3.scaffold_6496 18604682 70 + 26866924 CGUCAAAUGCAUUUGAUUAGCUGAAUUUUCGGCGUCGCUUGCCGCACUUCAAGUGCCACUGAC--GAUCCUU------- (((((...((((((((...(((((....)))))...((.....))...))))))))...))))--)......------- ( -23.20, z-score = -3.05, R) >droGri2.scaffold_15245 16314494 70 - 18325388 CGUCAACUCCAUUUGAUUAGCUGAAUUUUCGGCGUCGCUUGUCUCACUUCAAGUGCCAAAAAA--GAUCCUU------- .(((((......)))))..(((((....)))))(.((((((........)))))).)......--.......------- ( -13.90, z-score = -1.33, R) >consensus CGUAAACUAUAUUUGAUUAGCUGAAAUUUCGGCGUCGCUUGUCUCACUUCAAGUGGCUAUCCU__GAUCCUUGGCAGGA ..............((((.(((((....)))))..((((((........))))))..........)))).......... (-10.81 = -11.07 + 0.26)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:16:38 2011