
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,683,295 – 7,683,377 |
| Length | 82 |
| Max. P | 0.974245 |

| Location | 7,683,295 – 7,683,377 |
|---|---|
| Length | 82 |
| Sequences | 11 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 86.96 |
| Shannon entropy | 0.24367 |
| G+C content | 0.28520 |
| Mean single sequence MFE | -16.45 |
| Consensus MFE | -14.54 |
| Energy contribution | -14.81 |
| Covariance contribution | 0.27 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.88 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.90 |
| SVM RNA-class probability | 0.974245 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
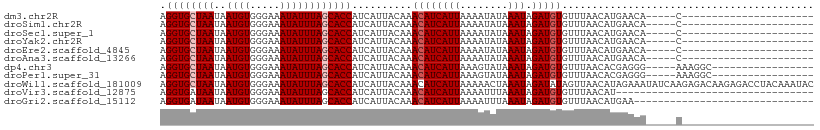
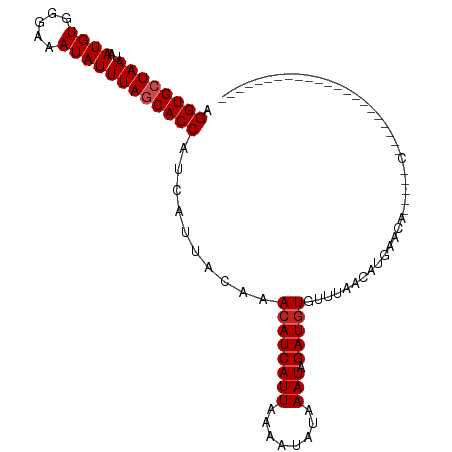
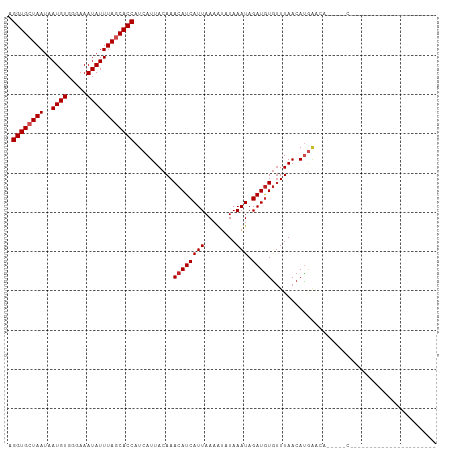
>dm3.chr2R 7683295 82 + 21146708 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA-----C---------------------- .((((((((..((((.....)))))))))))).((((...(((((.((((............)))))))))...))))...-----.---------------------- ( -18.30, z-score = -2.88, R) >droSim1.chr2R 6211060 82 + 19596830 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA-----C---------------------- .((((((((..((((.....)))))))))))).((((...(((((.((((............)))))))))...))))...-----.---------------------- ( -18.30, z-score = -2.88, R) >droSec1.super_1 5235307 82 + 14215200 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA-----C---------------------- .((((((((..((((.....)))))))))))).((((...(((((.((((............)))))))))...))))...-----.---------------------- ( -18.30, z-score = -2.88, R) >droYak2.chr2R 11868817 82 - 21139217 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA-----C---------------------- .((((((((..((((.....)))))))))))).((((...(((((.((((............)))))))))...))))...-----.---------------------- ( -18.30, z-score = -2.88, R) >droEre2.scaffold_4845 4476019 82 - 22589142 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA-----C---------------------- .((((((((..((((.....)))))))))))).((((...(((((.((((............)))))))))...))))...-----.---------------------- ( -18.30, z-score = -2.88, R) >droAna3.scaffold_13266 6661773 82 - 19884421 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA-----C---------------------- .((((((((..((((.....)))))))))))).((((...(((((.((((............)))))))))...))))...-----.---------------------- ( -18.30, z-score = -2.88, R) >dp4.chr3 16945919 87 + 19779522 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAGUAUAAAUAGAUGUGUUUAACACGAGGG-----AAAGGC----------------- .((((((((..((((.....))))))))))))..........((((((((........))).)))))..............-----......----------------- ( -16.20, z-score = -1.57, R) >droPer1.super_31 121293 87 - 935084 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAGUAUAAAUAGAUGUGUUUAACACGAGGG-----AAAGGC----------------- .((((((((..((((.....))))))))))))..........((((((((........))).)))))..............-----......----------------- ( -16.20, z-score = -1.57, R) >droWil1.scaffold_181009 3301503 109 - 3585778 AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAACUAAAUAGAUAUAGUUAACAUAGAAAUAUCAAGAGACAAGAGACCUACAAAUAC .((((((((..((((.....))))))))))))..............................(((((..(((...)))..)))))........................ ( -14.70, z-score = -1.43, R) >droVir3.scaffold_12875 15416010 76 - 20611582 AGGUGAUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUUUAAAUAGAUGUGUUUAACAU--------------------------------- .((((.....(((((.....)))))...))))..........((((((((........))).))))).........--------------------------------- ( -11.00, z-score = -0.60, R) >droGri2.scaffold_15112 4961075 79 - 5172618 AGGUGAUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUUUAAAUAGAUGUGUUUAACAUGAA------------------------------ .((((.....(((((.....)))))...)))).((((...(((((.((((............)))))))))...)))).------------------------------ ( -13.10, z-score = -0.99, R) >consensus AGGUGCUAAUAAUGUGGGAAAUAUUUAGCACCAUCAUUACAAACAUCAUUAAAAUAUAAAUAGAUGUGUUUAACAUGAACA_____C______________________ .((((((((..((((.....))))))))))))..........((((((((........))).))))).......................................... (-14.54 = -14.81 + 0.27)
| Location | 7,683,295 – 7,683,377 |
|---|---|
| Length | 82 |
| Sequences | 11 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 86.96 |
| Shannon entropy | 0.24367 |
| G+C content | 0.28520 |
| Mean single sequence MFE | -14.90 |
| Consensus MFE | -13.65 |
| Energy contribution | -13.75 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.39 |
| SVM RNA-class probability | 0.935223 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
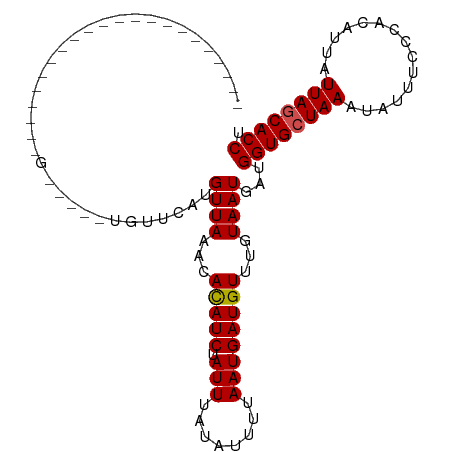
>dm3.chr2R 7683295 82 - 21146708 ----------------------G-----UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ----------------------.-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -16.06, z-score = -1.85, R) >droSim1.chr2R 6211060 82 - 19596830 ----------------------G-----UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ----------------------.-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -16.06, z-score = -1.85, R) >droSec1.super_1 5235307 82 - 14215200 ----------------------G-----UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ----------------------.-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -16.06, z-score = -1.85, R) >droYak2.chr2R 11868817 82 + 21139217 ----------------------G-----UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ----------------------.-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -16.06, z-score = -1.85, R) >droEre2.scaffold_4845 4476019 82 + 22589142 ----------------------G-----UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ----------------------.-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -16.06, z-score = -1.85, R) >droAna3.scaffold_13266 6661773 82 + 19884421 ----------------------G-----UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ----------------------.-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -16.06, z-score = -1.85, R) >dp4.chr3 16945919 87 - 19779522 -----------------GCCUUU-----CCCUCGUGUUAAACACAUCUAUUUAUACUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU -----------------......-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -15.56, z-score = -2.57, R) >droPer1.super_31 121293 87 + 935084 -----------------GCCUUU-----CCCUCGUGUUAAACACAUCUAUUUAUACUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU -----------------......-----...((((..(((((((((..(........)..))).))))))..)))).((((((((...............)))))))). ( -15.56, z-score = -2.57, R) >droWil1.scaffold_181009 3301503 109 + 3585778 GUAUUUGUAGGUCUCUUGUCUCUUGAUAUUUCUAUGUUAACUAUAUCUAUUUAGUUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ......(((((.....((((....))))..)))))((((...(((((.(((........))))))))...))))...((((((((...............)))))))). ( -17.86, z-score = -1.04, R) >droVir3.scaffold_12875 15416010 76 + 20611582 ---------------------------------AUGUUAAACACAUCUAUUUAAAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAUCACCU ---------------------------------..((((...(((((.(((........))))))))...))))...((((...((((..........))))..)))). ( -8.70, z-score = -0.07, R) >droGri2.scaffold_15112 4961075 79 + 5172618 ------------------------------UUCAUGUUAAACACAUCUAUUUAAAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAUCACCU ------------------------------.((((..(((((((((..(........)..))).))))))..)))).((((...((((..........))))..)))). ( -9.90, z-score = -0.42, R) >consensus ______________________G_____UGUUCAUGUUAAACACAUCUAUUUAUAUUUUAAUGAUGUUUGUAAUGAUGGUGCUAAAUAUUUCCCACAUUAUUAGCACCU ...................................((((...(((((.(((........))))))))...))))...((((((((...............)))))))). (-13.65 = -13.75 + 0.10)

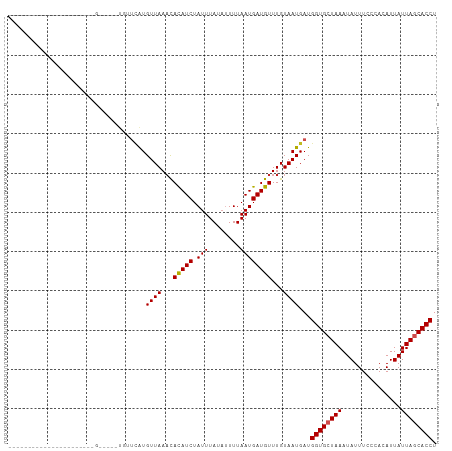
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:16:13 2011