
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,672,907 – 7,672,998 |
| Length | 91 |
| Max. P | 0.711891 |

| Location | 7,672,907 – 7,672,998 |
|---|---|
| Length | 91 |
| Sequences | 7 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 72.31 |
| Shannon entropy | 0.54253 |
| G+C content | 0.39978 |
| Mean single sequence MFE | -20.76 |
| Consensus MFE | -11.34 |
| Energy contribution | -11.86 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.47 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.711891 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
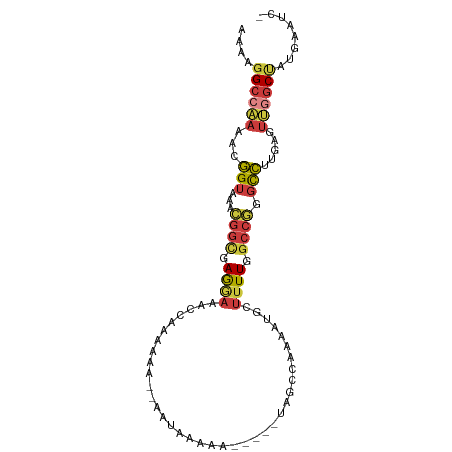
>dm3.chr2R 7672907 91 + 21146708 AAAAGGCCAAAACGGUAAACGGUGAGGAAACCAAAAAAUAAAUAAAAA-----UAGCCAAAAUGCUUUUGGCCGGGCCUUGAGUUGGCUAUGAAUC- ....((((((...(((...((.((.(....)))...............-----..((((((.....)))))))).))).....)))))).......- ( -21.60, z-score = -1.29, R) >droSim1.chr2R 6200316 92 + 19596830 AAAAGGCCAAAACGGUAAACGGCGAGGAAACCAAAAAAUAAGUAAAAA-----UAGCCAAAAUGCUUUUGGCCGGGUCUUGAGUUGGCUAUGAAAUC ....((((((..(((.(..(((((.(....))..............((-----.(((......))).)).))))..).)))..))))))........ ( -21.00, z-score = -1.10, R) >droSec1.super_1 5225032 92 + 14215200 AAAAGGCCAAAACGGUAAACGGCGAGGAAACCAAAAAAUAAAUAAAAA-----UAGCCAAAAUGCUUUUGGCCGGGCCUUGAGUUGGCUAUGAAAUC ....((((((...(((...(((((.(....))..............((-----.(((......))).)).)))).))).....))))))........ ( -22.30, z-score = -1.33, R) >droYak2.chr2R 11858116 89 - 21139217 AAAAGGCCAAAACGGUAAACGGCGAGGAAACCAAAAGA--AAUAAAAA-----UAGCCAAAAUGCUUUUGGCCGGGCCUUGAGUUGGCUAUGAAUC- ....((((((...(((...((((..(....)((((((.--........-----...........)))))))))).))).....)))))).......- ( -23.55, z-score = -1.61, R) >droEre2.scaffold_4845 4465736 89 - 22589142 AAAAGGCCAAAACGGUAAACGGCUAGGAAACCAAAAGA--AAUAAAAA-----UAGCCAAAAUGCUUUUGGCCGGGCCUUGAGUUGGCUAUGAAUC- ....(((((((..((((...((((((....)....(..--..).....-----)))))....))))))))))).((((.......)))).......- ( -24.60, z-score = -1.96, R) >droAna3.scaffold_13266 6652227 81 - 19884421 AAAAUGCCAAA-CGGUAAAUGGUAAAAAAAAUCACACA---AAAAAAA-----UAC------UGUUUUUGGCCAGGCCUUGAGGCCAC-ACGGUUUU .....(((...-.(((...((((.......))))..((---((((...-----...------..))))))))).((((....))))..-..)))... ( -16.90, z-score = -0.52, R) >droMoj3.scaffold_6496 24885986 96 + 26866924 AAAAGGCCGAAACUGCCAAAAAGAAAAAAUAUAAAUAAGUAAAAAAAAGCUAACAAUUUUCGUGGGCUUGGCCACACAAAACGUCAGCCGCAGUGG- ...........(((((..............................(((((.((.......)).)))))(((...((.....))..))))))))..- ( -15.40, z-score = 0.33, R) >consensus AAAAGGCCAAAACGGUAAACGGCGAGGAAACCAAAAAA__AAUAAAAA_____UAGCCAAAAUGCUUUUGGCCGGGCCUUGAGUUGGCUAUGAAUC_ ....((((((...(((...((((.((((.....................................)))).)))).))).....))))))........ (-11.34 = -11.86 + 0.52)


| Location | 7,672,907 – 7,672,998 |
|---|---|
| Length | 91 |
| Sequences | 7 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 72.31 |
| Shannon entropy | 0.54253 |
| G+C content | 0.39978 |
| Mean single sequence MFE | -19.52 |
| Consensus MFE | -9.25 |
| Energy contribution | -10.54 |
| Covariance contribution | 1.29 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.20 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.509513 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 7672907 91 - 21146708 -GAUUCAUAGCCAACUCAAGGCCCGGCCAAAAGCAUUUUGGCUA-----UUUUUAUUUAUUUUUUGGUUUCCUCACCGUUUACCGUUUUGGCCUUUU -................((((((.(((((((.....))))))).-----................(((......)))............)))))).. ( -18.90, z-score = -1.26, R) >droSim1.chr2R 6200316 92 - 19596830 GAUUUCAUAGCCAACUCAAGACCCGGCCAAAAGCAUUUUGGCUA-----UUUUUACUUAUUUUUUGGUUUCCUCGCCGUUUACCGUUUUGGCCUUUU .........(((((..(.......(((((((.....))))))).-----................(((......))).......)..)))))..... ( -17.40, z-score = -1.03, R) >droSec1.super_1 5225032 92 - 14215200 GAUUUCAUAGCCAACUCAAGGCCCGGCCAAAAGCAUUUUGGCUA-----UUUUUAUUUAUUUUUUGGUUUCCUCGCCGUUUACCGUUUUGGCCUUUU .................((((((.(((((((.....))))))).-----................(((......)))............)))))).. ( -19.20, z-score = -0.94, R) >droYak2.chr2R 11858116 89 + 21139217 -GAUUCAUAGCCAACUCAAGGCCCGGCCAAAAGCAUUUUGGCUA-----UUUUUAUU--UCUUUUGGUUUCCUCGCCGUUUACCGUUUUGGCCUUUU -................((((((.(((((((.....))))))).-----........--......(((......)))............)))))).. ( -19.20, z-score = -0.90, R) >droEre2.scaffold_4845 4465736 89 + 22589142 -GAUUCAUAGCCAACUCAAGGCCCGGCCAAAAGCAUUUUGGCUA-----UUUUUAUU--UCUUUUGGUUUCCUAGCCGUUUACCGUUUUGGCCUUUU -........(((.......)))..((((((((......((((((-----.....(((--......)))....)))))).......)))))))).... ( -20.82, z-score = -1.39, R) >droAna3.scaffold_13266 6652227 81 + 19884421 AAAACCGU-GUGGCCUCAAGGCCUGGCCAAAAACA------GUA-----UUUUUUU---UGUGUGAUUUUUUUUACCAUUUACCG-UUUGGCAUUUU ........-..((((....))))..((((((....------(((-----.......---((.((((......))))))..)))..-))))))..... ( -17.40, z-score = -0.64, R) >droMoj3.scaffold_6496 24885986 96 - 26866924 -CCACUGCGGCUGACGUUUUGUGUGGCCAAGCCCACGAAAAUUGUUAGCUUUUUUUUACUUAUUUAUAUUUUUUCUUUUUGGCAGUUUCGGCCUUUU -((((((((((((((.(((((((.(((...))))))))))...)))))))........................(.....))))))...))...... ( -23.70, z-score = -2.28, R) >consensus _AAUUCAUAGCCAACUCAAGGCCCGGCCAAAAGCAUUUUGGCUA_____UUUUUAUU__UUUUUUGGUUUCCUCGCCGUUUACCGUUUUGGCCUUUU .........(((.......)))..((((((((((......)))......................(((......))).........))))))).... ( -9.25 = -10.54 + 1.29)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:16:11 2011