
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,479,250 – 7,479,355 |
| Length | 105 |
| Max. P | 0.640877 |

| Location | 7,479,250 – 7,479,355 |
|---|---|
| Length | 105 |
| Sequences | 12 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 74.78 |
| Shannon entropy | 0.52846 |
| G+C content | 0.66834 |
| Mean single sequence MFE | -51.58 |
| Consensus MFE | -22.69 |
| Energy contribution | -22.78 |
| Covariance contribution | 0.09 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.640877 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
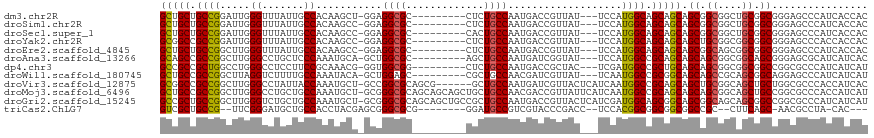
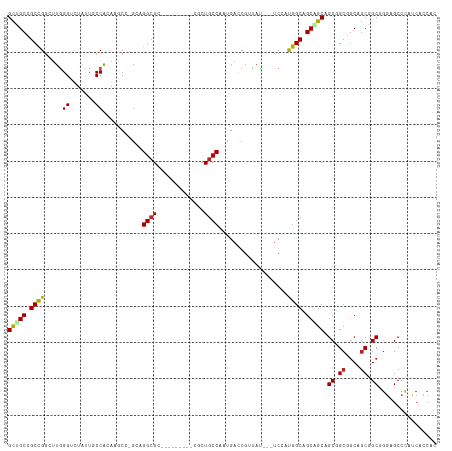
>dm3.chr2R 7479250 105 - 21146708 GCUGCUGCCGGAUUGGGUUUAUUGCCACAAGCU-GGAGGCGC---------CUCUGCCAAUGACCGUUAU---UCCAUGGCAGCAGCAGCGGCGGCUGCGGCGGGAGCCCAUCACCAC (((.(((((((.((((((.....))).))).))-(.((.(((---------(.((((...((.((((...---...))))))...)))).)))).)).)))))).))).......... ( -44.40, z-score = -0.46, R) >droSim1.chr2R 6003763 105 - 19596830 GCUGCUGCCGGAUUGGGUUUAUUGCCACAAGCC-GGAGGCGC---------CUCUGCCAAUGACCGUUAU---UCCAUGGCAGCAGCAGCGGCGGCUGCGGCGGGAGCCCAUCACCAC (((.(((((((.((((((.....))).))).))-(.((.(((---------(.((((...((.((((...---...))))))...)))).)))).)).)))))).))).......... ( -47.40, z-score = -1.28, R) >droSec1.super_1 5030070 105 - 14215200 GCUGCUGCCGGAUUGGGUUUAUUGCCACAAGCC-GGAGGCGC---------CACUGCCAAUGACCGUUAU---UCCAUGGCAGCAGCAGCGGCGGCUGCGGCGGGAGCCCAUCACCAC (((.(((((((.((((((.....))).))).))-(.((.(((---------(.((((...((.((((...---...))))))...)))).)))).)).)))))).))).......... ( -47.80, z-score = -1.52, R) >droYak2.chr2R 11663155 105 + 21139217 GCGGCCGCCGGAUUGGGUUUAUUGCCACAAGCC-GGAGGCGC---------CUCUGCCAAUGACCGUUAU---UCCAUGGCAGCAGCAGCUGCGGCGGCGGCGGGAGCCCACCACCAC ...(((.((((.((((((.....))).))).))-)).)))((---------(.(((((......(((...---...)))(((((....)))))))))).)))((....))........ ( -46.50, z-score = -0.71, R) >droEre2.scaffold_4845 4272256 105 + 22589142 GCUGCUGCCGGCUUGGGUUUAUUGCCACAAGCC-GGAGGCGC---------CUCUGCCAAUGACCGUUAU---UCCAUGGCAGCAGCAGCGGCAGCGGCGGCGGGAGCCCAUCACCAC ((((((((((((((((((.....))).))))))-....((((---------..((((((.((........---..))))))))..)).)))))))))))(((....)))......... ( -53.90, z-score = -2.82, R) >droAna3.scaffold_13266 1193482 105 - 19884421 GCAGCCGCCGGCUUGGGCCUGCUCCCAAAUGCA-GCUGGCGC---------AGCUGCCAAUGAUCGGUAU---UCCAUGGCCGCAGCAGCAGCGGCGGCAGCGGGAGCGCAUCAUCAC (((((((((((((((((......)))).....)-))))))).---------.)))))...((((..(..(---(((...(((((.((....)).)))))....))))..)...)))). ( -48.40, z-score = -0.22, R) >dp4.chr3 4666837 105 - 19779522 GCCGCCGCUGGCCUGGGCCUCCUUCCGCAAACG-GGUGGCGG---------CUCUGCCAAUGACCGCUAC---UCGAUGGCCGCUGCAGCAGCGGCGGCGGCCGGCGCCCAUCAUCAU (((((((((((((.(((......))).....((-((((((((---------.((.......)))))))))---)))..))))((....)))))))))))(((....)))......... ( -54.40, z-score = -1.48, R) >droWil1.scaffold_180745 960129 105 + 2843958 GCUGCCGCCGGCUUAGGUCUUUUGCCAAAUACA-GCUGGAGC---------CGCUGCCAACGAUCGUUAU---UCAAUGGCCGCGGCAGCAGCCGCAGCGGCAGGAGCCCAUCAUCAU .((((((((((((..((.(((..((........-))..))))---------)((((((..((..((((..---..))))..)).)))))))))))..))))))).............. ( -45.10, z-score = -2.00, R) >droVir3.scaffold_12875 7642687 111 + 20611582 GCGGCCGCCGGCUUGGGCCUAUUACCAAAUGCU-GCCGGCGCAGCG------GCUGCCAAUGAUCGUUACUCAUCAAUGGCCGCAGCAGCUGCGGCAGCUGCUGGCGCCCACCAUCAC ..((.(((((((((((........))))..(((-(((...(((((.------((.((((.((((........)))).)))).))....))))))))))).))))))).))........ ( -53.40, z-score = -1.78, R) >droMoj3.scaffold_6496 6633184 117 + 26866924 GCUGCCGCCGGCUUGGGCCUGCUGCCAAAUGCU-GCGGGCGCAGCAGCAGCUGCUGCCAACGACCGUUAUUCAUCAAUGGCCGCAGCAGCAGCGGCAGCUGCCGGCGCCCACCAUCAU .....(((((((....(((.(((((....((((-(((((.(((((((...)))))))).....(((((.......)))))))))))))))))))))....)))))))........... ( -59.30, z-score = -1.28, R) >droGri2.scaffold_15245 7398039 117 - 18325388 GCCGCUGCCGGCUUGGGUCUGCUGCCAAAUGCU-GCGGGCGCAGCAGCUGCCGCUGCCAAUGACCGUUACUCAUCGAUGGCAGCGGCAGCGGCAGCAGCGGCCGGCGCCCAUCAUCAU ...((.(((((((...(.((((((((...((((-((....))))))(((((((((((((((((.......))))...))))))))))))))))))))))))))))))).......... ( -73.60, z-score = -4.60, R) >triCas2.ChLG7 6229712 99 + 17478683 GUCGCUGCCG--UUCGGGAUGCUGCCACCUACGAGCGGGCGCG--------GGAUGCCGUCGUACCCGACC--UCCACGGCGGCGGCGGCCGC--CUUCAGC-AACGCCUA-CAC--- ((((((((((--((((((.(((.(((..(.....)..)))(((--------(....)))).))))))))..--...)))))))))))(((.((--.....))-...)))..-...--- ( -44.80, z-score = -0.39, R) >consensus GCUGCCGCCGGCUUGGGUCUAUUGCCACAAGCC_GCAGGCGC_________CGCUGCCAAUGACCGUUAU___UCCAUGGCAGCAGCAGCGGCGGCAGCGGCGGGAGCCCAUCACCAC (((((.((((.....((.......))...........((((.............))))...................)))).))))).((.((....)).))................ (-22.69 = -22.78 + 0.09)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:15:39 2011