
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,356,792 – 7,356,892 |
| Length | 100 |
| Max. P | 0.997582 |

| Location | 7,356,792 – 7,356,892 |
|---|---|
| Length | 100 |
| Sequences | 10 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 82.89 |
| Shannon entropy | 0.32990 |
| G+C content | 0.39990 |
| Mean single sequence MFE | -28.23 |
| Consensus MFE | -15.93 |
| Energy contribution | -16.73 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.901323 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
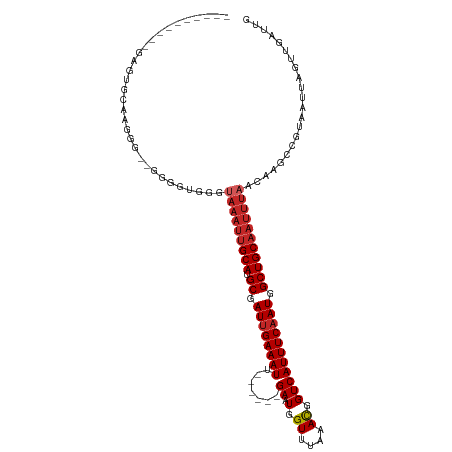
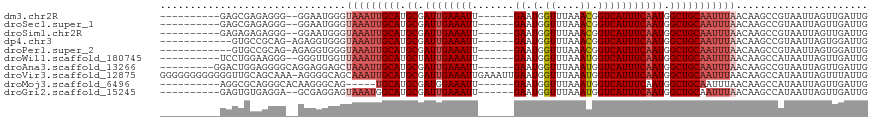
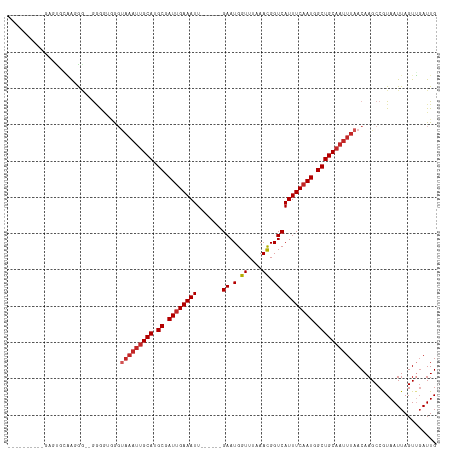
>dm3.chr2R 7356792 100 + 21146708 ----------GAGCGAGAGGG--GGAAUGGGUAAAUUGCAUGCGAUUGAAAUU------GAAUGGUUUAAACGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUUGAUUG ----------.(((.....(.--((..((..(((((((((.((.((((((((.------((.((.......)).)))))))))).))))))))))).))..)).).....)))..... ( -28.90, z-score = -3.31, R) >droSec1.super_1 4909345 100 + 14215200 ----------GAGCGAGAGGG--GGAAUGGGUAAAUUGCAUGCGAUUGAAAUU------GAAUGGUUUAAACGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUUGAUUG ----------.(((.....(.--((..((..(((((((((.((.((((((((.------((.((.......)).)))))))))).))))))))))).))..)).).....)))..... ( -28.90, z-score = -3.31, R) >droSim1.chr2R 5883009 100 + 19596830 ----------GAGAGAGAGGG--GGAAUGGGUAAAUUGCAUGCGAUUGAAAUU------GAAUGGUUUAAACGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUUGAUUG ----------...........--((..((..(((((((((.((.((((((((.------((.((.......)).)))))))))).))))))))))).))..))............... ( -26.90, z-score = -3.15, R) >dp4.chr3 4539147 99 + 19779522 ------------GUGCCGCAG-AGAGGUGGGUAAAUUGCAUGCGAUUGAAAUU------GAAUGGUUUAAACGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUGGAUUG ------------...((((.(-(..(((..((((((((((.((.((((((((.------((.((.......)).)))))))))).)))))))))).))..)))....)).)))).... ( -33.10, z-score = -3.66, R) >droPer1.super_2 4728064 99 + 9036312 ------------GUGCCGCAG-AGAGGUGGGUAAAUUGCAUGCGAUUGAAAUU------GAAUGGUUUAAACGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUGGAUUG ------------...((((.(-(..(((..((((((((((.((.((((((((.------((.((.......)).)))))))))).)))))))))).))..)))....)).)))).... ( -33.10, z-score = -3.66, R) >droWil1.scaffold_180745 766819 100 - 2843958 ----------UCCUGGAAGGG--GGGUUGGUUAAAUUGCAUGCUAUUGAAAUU------GAAUGGUUUAAAUGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCAUAAUUAGUUGAUUG ----------((((....)))--)((((.(((((((((((.(((((((((((.------((.(.((....)).))))))))))))))))))))))))).))))............... ( -34.20, z-score = -4.54, R) >droAna3.scaffold_13266 1072292 103 + 19884421 ---------GGACUGGAGGGGCAGGAGGAGCUAAAUUGCAUGCGAUUGAAAUU------GAAUGGUUUAAAUGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUUGAUUG ---------.((((((.(.(((.(......)(((((((((.((.((((((((.------((.(.((....)).))))))))))).)))))))))))....))).)..))))))..... ( -26.20, z-score = -1.62, R) >droVir3.scaffold_12875 7487703 117 - 20611582 GGGGGGGGGGGGUUGCAGCAAA-AGGGGCAGCAAAUUGCAUGCGAUUGAAAUUGAAAUUGAAUGGUUUAAAUGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCAUAAUUAGUUUAUUG .((...(..((((((((((...-....(((((.....)).))).((((((((.((.(((..........)))..)))))))))).))))))))))..)...))............... ( -29.30, z-score = -1.61, R) >droMoj3.scaffold_6496 6463488 97 - 26866924 ----------AGGCGCAGGGCACAAGGGCAG-----UGCAUGCGAUGGAAAUU------GAAUGGUUUAAAUGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCAUAAUUAGUUGAUUG ----------...((((..((((.......)-----))).)))).....((((------(((((((((((((((((((....))))))...))))))...)))))..))))))..... ( -22.90, z-score = -0.07, R) >droGri2.scaffold_15245 7236968 100 + 18325388 ----------GAGUGUGAGGA--GCGAGGAGUAAAUGGCAUGCGAUUGAAAUU------GAAUGGUUUAAAUGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCAUAAUUAGUUGAUUG ----------...((((....--((..(...(((((.(((.((.((((((((.------((.(.((....)).))))))))))).))))).))))).)..))))))............ ( -18.80, z-score = 0.38, R) >consensus __________GAGUGCAAGGG__GGGGUGGGUAAAUUGCAUGCGAUUGAAAUU______GAAUGGUUUAAACGGUCAUUUCAAUGGCUGCAAUUUAACAAGCCGUAAUUAGUUGAUUG ...............................(((((((((.((.((((((((........((....))........)))))))).)))))))))))...................... (-15.93 = -16.73 + 0.80)

| Location | 7,356,792 – 7,356,892 |
|---|---|
| Length | 100 |
| Sequences | 10 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 82.89 |
| Shannon entropy | 0.32990 |
| G+C content | 0.39990 |
| Mean single sequence MFE | -19.93 |
| Consensus MFE | -15.09 |
| Energy contribution | -15.71 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.05 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.13 |
| SVM RNA-class probability | 0.997582 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
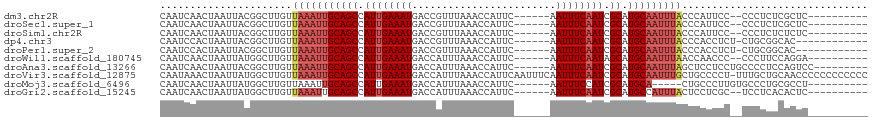
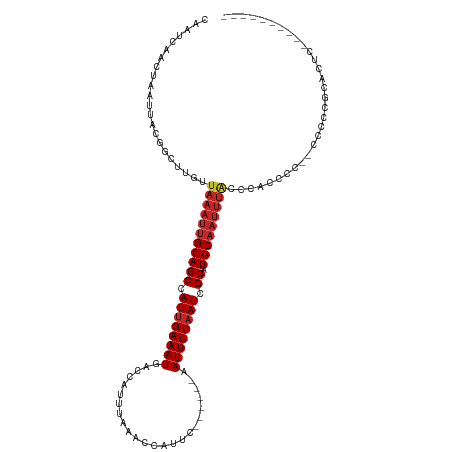
>dm3.chr2R 7356792 100 - 21146708 CAAUCAACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCGUUUAAACCAUUC------AAUUUCAAUCGCAUGCAAUUUACCCAUUCC--CCCUCUCGCUC---------- ...............((..((.(((((((((((.((((((((((..((....))...))------.)))))))).)).)))))))))..))..))--...........---------- ( -20.00, z-score = -4.02, R) >droSec1.super_1 4909345 100 - 14215200 CAAUCAACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCGUUUAAACCAUUC------AAUUUCAAUCGCAUGCAAUUUACCCAUUCC--CCCUCUCGCUC---------- ...............((..((.(((((((((((.((((((((((..((....))...))------.)))))))).)).)))))))))..))..))--...........---------- ( -20.00, z-score = -4.02, R) >droSim1.chr2R 5883009 100 - 19596830 CAAUCAACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCGUUUAAACCAUUC------AAUUUCAAUCGCAUGCAAUUUACCCAUUCC--CCCUCUCUCUC---------- ...............((..((.(((((((((((.((((((((((..((....))...))------.)))))))).)).)))))))))..))..))--...........---------- ( -20.00, z-score = -4.81, R) >dp4.chr3 4539147 99 - 19779522 CAAUCCACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCGUUUAAACCAUUC------AAUUUCAAUCGCAUGCAAUUUACCCACCUCU-CUGCGGCAC------------ (((.((.........)).))).(((((((((((.((((((((((..((....))...))------.)))))))).)).)))))))))((((.....-.)).))...------------ ( -19.00, z-score = -2.10, R) >droPer1.super_2 4728064 99 - 9036312 CAAUCCACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCGUUUAAACCAUUC------AAUUUCAAUCGCAUGCAAUUUACCCACCUCU-CUGCGGCAC------------ (((.((.........)).))).(((((((((((.((((((((((..((....))...))------.)))))))).)).)))))))))((((.....-.)).))...------------ ( -19.00, z-score = -2.10, R) >droWil1.scaffold_180745 766819 100 + 2843958 CAAUCAACUAAUUAUGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCAUUUAAACCAUUC------AAUUUCAAUAGCAUGCAAUUUAACCAACCC--CCCUUCCAGGA---------- ...............((.(((((((((((((((.((((((((.................------.)))))))).)).)))))))))).))).))--.((.....)).---------- ( -23.07, z-score = -4.07, R) >droAna3.scaffold_13266 1072292 103 - 19884421 CAAUCAACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCAUUUAAACCAUUC------AAUUUCAAUCGCAUGCAAUUUAGCUCCUCCUGCCCCUCCAGUCC--------- ...............(((..(((((((((((((.((((((((.................------.)))))))).)).))))))))))).......)))..........--------- ( -24.37, z-score = -4.79, R) >droVir3.scaffold_12875 7487703 117 + 20611582 CAAUAAACUAAUUAUGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCAUUUAAACCAUUCAAUUUCAAUUUCAAUCGCAUGCAAUUUGCUGCCCCU-UUUGCUGCAACCCCCCCCCCCC ...............(((..((.((((((((((.((((((((........................)))))))).)).)))))))))).)))...-...................... ( -21.36, z-score = -2.51, R) >droMoj3.scaffold_6496 6463488 97 + 26866924 CAAUCAACUAAUUAUGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCAUUUAAACCAUUC------AAUUUCCAUCGCAUGCA-----CUGCCCUUGUGCCCUGCGCCU---------- ...(((.......((((((.((......)))))))).....)))...............------.........((((.(((-----(.......))))..))))...---------- ( -17.80, z-score = -0.86, R) >droGri2.scaffold_15245 7236968 100 - 18325388 CAAUCAACUAAUUAUGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCAUUUAAACCAUUC------AAUUUCAAUCGCAUGCCAUUUACUCCUCGC--UCCUCACACUC---------- .............(((((.(((......)))((.((((((((.................------.)))))))).))..)))))...........--...........---------- ( -14.67, z-score = -2.06, R) >consensus CAAUCAACUAAUUACGGCUUGUUAAAUUGCAGCCAUUGAAAUGACCAUUUAAACCAUUC______AAUUUCAAUCGCAUGCAAUUUACCCACCCC__CCCCCGCACUC__________ ......................(((((((((((.((((((((........................)))))))).)).)))))))))............................... (-15.09 = -15.71 + 0.62)
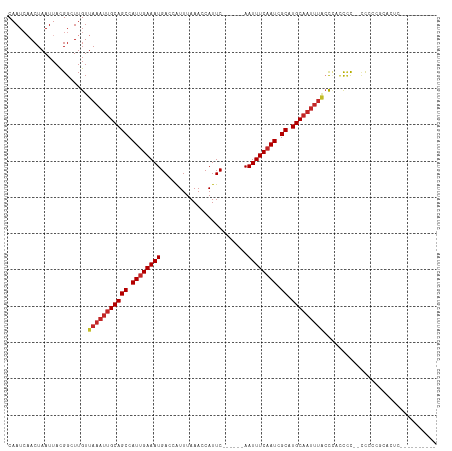
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:15:22 2011