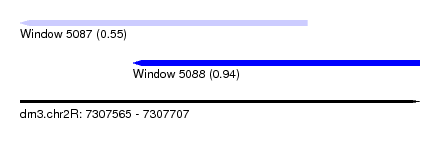
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 7,307,565 – 7,307,707 |
| Length | 142 |
| Max. P | 0.938715 |

| Location | 7,307,565 – 7,307,667 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 61.14 |
| Shannon entropy | 0.62248 |
| G+C content | 0.43576 |
| Mean single sequence MFE | -21.14 |
| Consensus MFE | -11.14 |
| Energy contribution | -11.10 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.59 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.546581 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
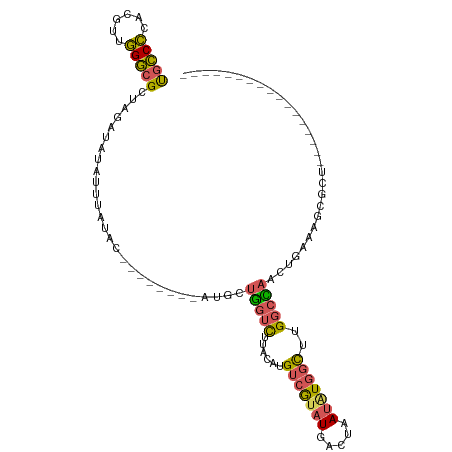
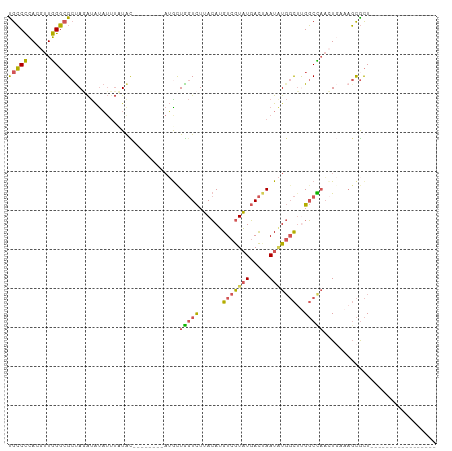
>dm3.chr2R 7307565 102 - 21146708 UGCCCCACGUUGGGCGCUAGAUAUAUUUCUAC--------AUGCUGGUCUUACAUGUCGUAUGACUAAUAUGGCUUGGCCAACUGAAAGCGCUCCCCUCUCGAAUUUAUG ...........((((((((((......))).(--------(.(.(((((......(((((((.....)))))))..))))).)))..)))))))................ ( -24.60, z-score = -0.80, R) >droAna3.scaffold_13266 1010740 82 - 19884421 UGGCUUAC--UAGUCG-UAUGCGCAAUAUUAU--------CAGCUACUUUAAAAAGUCAUUUUAUUAAUCUCCUUCGUCUUUUUCCCUCUAUU----------------- ((((((..--(((..(-((.((..........--------..))))).)))..))))))..................................----------------- ( -6.70, z-score = -0.18, R) >droSim1.chr2R 5833306 85 - 19596830 UGCCCCACGUUGGGCGCUAGAUAUAUUUCUGC--------AUGUUGGUCUUACAUGUCGUAUGACUAAUAUGGCUUGGCCAACUGAAAGCGCU----------------- .((((......))))((((((......))).(--------(.(((((((......(((((((.....)))))))..)))))))))..)))...----------------- ( -27.40, z-score = -1.92, R) >droSec1.super_1 4860700 85 - 14215200 UGCCCCACGUUGGGCGCUAGAUAUAUUUAUGC--------AUACUGGUCUUACAUGUCGUAUGACUAAUAUGGCUUGGCCAACUGAAAGCGCU----------------- .((((......))))((..............(--------(((.(((((.(((.....))).))))).))))((((..........)))))).----------------- ( -22.20, z-score = -0.39, R) >droYak2.chr2R 11483658 109 + 21139217 CGCCCCACGUUGGGAGCCAUAUAUUUGUAUAUUGAAUAAGAGGCUGGUCUUACAUGUCGUAUGACGAAUGUGGCACGGCUAAAU-AAAGCGCUCCUCUCUCGAAUUUAUG .......((..(((((((...((((((.....))))))...((((((((..((((.(((.....)))))))))).)))))....-...).))))))....))........ ( -24.80, z-score = 0.32, R) >consensus UGCCCCACGUUGGGCGCUAGAUAUAUUUAUAC________AUGCUGGUCUUACAUGUCGUAUGACUAAUAUGGCUUGGCCAACUGAAAGCGCU_________________ .((((......)))).............................(((((......(((((((.....)))))))..)))))............................. (-11.14 = -11.10 + -0.04)

| Location | 7,307,605 – 7,307,707 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 70.59 |
| Shannon entropy | 0.47852 |
| G+C content | 0.36339 |
| Mean single sequence MFE | -22.58 |
| Consensus MFE | -13.41 |
| Energy contribution | -12.35 |
| Covariance contribution | -1.06 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.938715 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
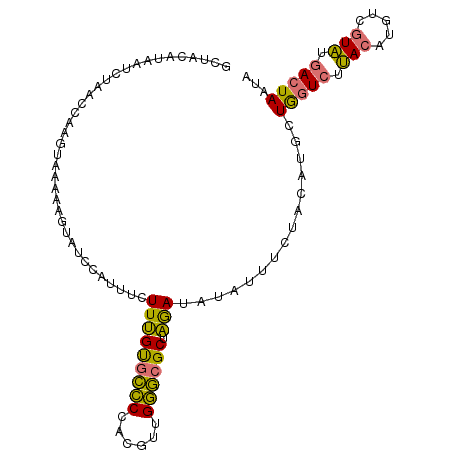
>dm3.chr2R 7307605 102 - 21146708 GCUACAUAAUCGAACCGAUUAAGAAGUAUCCAUUUCUUUGUGCCCCACGUUGGGCGCUAGAUAUAUUUCUACAUGCUGGUCUUACAUGUCGUAUGACUAAUA ((....((((((...))))))((((((((.....(((..((((((......)))))).))).))))))))....))(((((.(((.....))).)))))... ( -28.00, z-score = -2.91, R) >droAna3.scaffold_13266 1010763 91 - 19884421 ----CACAGAUUAUCAAAGAAAUGACUA-CUAUGUUUUGGCUUACUA-GUCGUAUGCGCAAUAU-----UAUCAGCUACUUUAAAAAGUCAUUUUAUUAAUC ----....(((((.....(((((((((.-.........((((....)-)))(((.((.......-----.....))))).......)))))))))..))))) ( -15.60, z-score = -1.29, R) >droSim1.chr2R 5833329 102 - 19596830 GCUACAUAAUCUAAGCAAGUAAAAAGUAUCCAUUUCUUUGUGCCCCACGUUGGGCGCUAGAUAUAUUUCUGCAUGUUGGUCUUACAUGUCGUAUGACUAAUA (((..........)))..(((..((((((.....(((..((((((......)))))).))).)))))).))).((((((((.(((.....))).)))))))) ( -24.30, z-score = -1.65, R) >droSec1.super_1 4860723 102 - 14215200 GCUACAUAAUCUAAGCAAGUAAAAAGUAUCCAUUUCUUUGUGCCCCACGUUGGGCGCUAGAUAUAUUUAUGCAUACUGGUCUUACAUGUCGUAUGACUAAUA (((..........)))........(((((.(((.(((..((((((......)))))).))).......))).)))))((((.(((.....))).)))).... ( -22.40, z-score = -1.08, R) >consensus GCUACAUAAUCUAACCAAGUAAAAAGUAUCCAUUUCUUUGUGCCCCACGUUGGGCGCUAGAUAUAUUUCUACAUGCUGGUCUUACAUGUCGUAUGACUAAUA ....................................(((((((((......)))))).)))...............(((((.(((.....))).)))))... (-13.41 = -12.35 + -1.06)

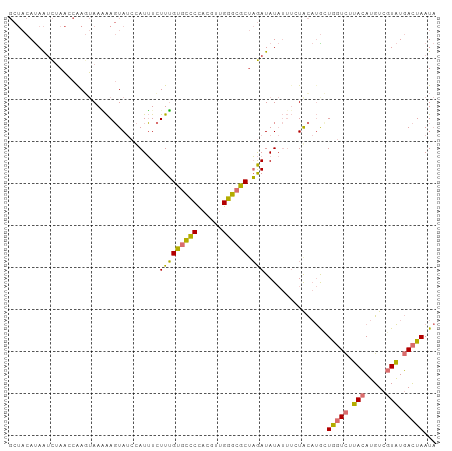
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:15:18 2011