
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 2,539,269 – 2,539,363 |
| Length | 94 |
| Max. P | 0.874692 |

| Location | 2,539,269 – 2,539,363 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 94.26 |
| Shannon entropy | 0.10094 |
| G+C content | 0.47234 |
| Mean single sequence MFE | -31.62 |
| Consensus MFE | -26.76 |
| Energy contribution | -27.52 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.719702 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 2539269 94 + 23011544 AGCCAAGAGUAUGGCAAUUUGUAGCUGGAAAAGUUUCGCUUCCCUGCAUGUCAGUUGAUUUUGAGGGAAUUGCGACAGAGUUCUCUGCUGUGGA .((((......))))..((..((((.((((.....((((((((((.((.(((....)))..))))))))..)))).....))))..))))..)) ( -31.10, z-score = -1.88, R) >droSim1.chr2L 2501672 94 + 22036055 AGCCAAGAGUAUGGCAAUUUGUAGCUGGAAAAGUUUCGCUUCCCGGCAAGUCAGUUGAUUUUGCGGGAAUUGCGACAAGGUUCUCUGCUGUGGA .((((......))))..((..((((.((((.....((((((((((.((((((....)).))))))))))..)))).....))))..))))..)) ( -32.80, z-score = -2.03, R) >droSec1.super_5 713214 94 + 5866729 AGCCAAGAGUAUGGCAAUUUGUAGCUGGAAAAGUUUCGCUUCCCGGCAAGUCAGUUGAUUUUGCGGGAAUUGCGACAGAGUUCUCUGCUGUGGA .((((......))))..((..((((.((((.....((((((((((.((((((....)).))))))))))..)))).....))))..))))..)) ( -32.80, z-score = -2.03, R) >droYak2.chr2L 2531854 94 + 22324452 AGCCAGGAGUAUGGCAAUUUGUAGCUGGCAAAGUUUCGCUUCCCAGCAAGUCAGUUGAUUUUGCGGGAAUUGCGACAGAGUUUUCGUCUGUGGA .((((......))))..((..(((.(((.(((.(((((((((((.(((((((....)).))))))))))..))))..)).)))))).)))..)) ( -28.70, z-score = -1.06, R) >droEre2.scaffold_4929 2585055 94 + 26641161 AGCCAGGAGUAUGGCAAUUUGUAGCUGGAAAAGUUUCGCUUCCCUGCAAGUCAGUUGAUUUUGCGGGAAUUGCGACCGAGUUUUCUUCUGUGGA .((((......))))..((..(((..((((((.(((((((((((.(((((((....)).))))))))))..))))..)).)))))).)))..)) ( -32.70, z-score = -2.35, R) >consensus AGCCAAGAGUAUGGCAAUUUGUAGCUGGAAAAGUUUCGCUUCCCGGCAAGUCAGUUGAUUUUGCGGGAAUUGCGACAGAGUUCUCUGCUGUGGA .((((......))))..((..((((.((((.....(((((((((.(((((((....)).))))))))))..)))).....))).).))))..)) (-26.76 = -27.52 + 0.76)


| Location | 2,539,269 – 2,539,363 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 94.26 |
| Shannon entropy | 0.10094 |
| G+C content | 0.47234 |
| Mean single sequence MFE | -24.09 |
| Consensus MFE | -19.86 |
| Energy contribution | -20.22 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.02 |
| SVM RNA-class probability | 0.874692 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 2539269 94 - 23011544 UCCACAGCAGAGAACUCUGUCGCAAUUCCCUCAAAAUCAACUGACAUGCAGGGAAGCGAAACUUUUCCAGCUACAAAUUGCCAUACUCUUGGCU .....(((.(((((.....((((..((((((((...((....))..)).))))))))))...)))))..))).......((((......)))). ( -23.70, z-score = -2.22, R) >droSim1.chr2L 2501672 94 - 22036055 UCCACAGCAGAGAACCUUGUCGCAAUUCCCGCAAAAUCAACUGACUUGCCGGGAAGCGAAACUUUUCCAGCUACAAAUUGCCAUACUCUUGGCU .....(((.(((((.....((((..(((((((((..((....)).))).))))))))))...)))))..))).......((((......)))). ( -25.80, z-score = -2.78, R) >droSec1.super_5 713214 94 - 5866729 UCCACAGCAGAGAACUCUGUCGCAAUUCCCGCAAAAUCAACUGACUUGCCGGGAAGCGAAACUUUUCCAGCUACAAAUUGCCAUACUCUUGGCU .....(((.(((((.....((((..(((((((((..((....)).))).))))))))))...)))))..))).......((((......)))). ( -25.80, z-score = -2.73, R) >droYak2.chr2L 2531854 94 - 22324452 UCCACAGACGAAAACUCUGUCGCAAUUCCCGCAAAAUCAACUGACUUGCUGGGAAGCGAAACUUUGCCAGCUACAAAUUGCCAUACUCCUGGCU ....((((.......))))((((..(((((((((..((....)).)))).)))))))))......(((((..................))))). ( -22.77, z-score = -2.26, R) >droEre2.scaffold_4929 2585055 94 - 26641161 UCCACAGAAGAAAACUCGGUCGCAAUUCCCGCAAAAUCAACUGACUUGCAGGGAAGCGAAACUUUUCCAGCUACAAAUUGCCAUACUCCUGGCU .....((..(((((.....((((..(((((((((..((....)).)))).)))))))))...)))))...)).......((((......)))). ( -22.40, z-score = -1.74, R) >consensus UCCACAGCAGAGAACUCUGUCGCAAUUCCCGCAAAAUCAACUGACUUGCAGGGAAGCGAAACUUUUCCAGCUACAAAUUGCCAUACUCUUGGCU .........(((((.....((((..(((((((((..((....)).)))).)))))))))...)))))............((((......)))). (-19.86 = -20.22 + 0.36)
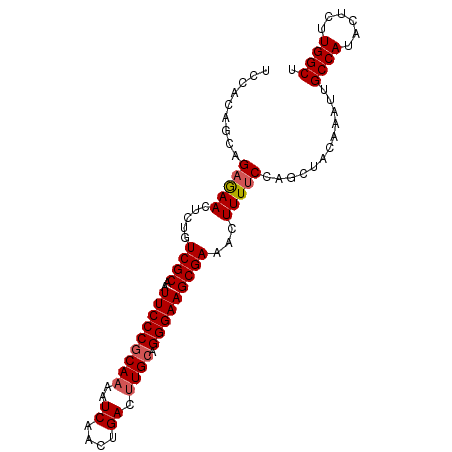
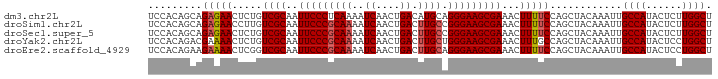



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:11:41 2011