
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,966,980 – 6,967,034 |
| Length | 54 |
| Max. P | 0.650586 |

| Location | 6,966,980 – 6,967,034 |
|---|---|
| Length | 54 |
| Sequences | 4 |
| Columns | 54 |
| Reading direction | forward |
| Mean pairwise identity | 85.19 |
| Shannon entropy | 0.24038 |
| G+C content | 0.29630 |
| Mean single sequence MFE | -7.30 |
| Consensus MFE | -5.60 |
| Energy contribution | -6.48 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.77 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.650586 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6966980 54 + 21146708 UUAGUUAAACUUUGCUAAGUAAAGUAUCGUAGCUUUUUCAUAGUUUCCGUCUUC ..(((((.(((((((...)))))))....))))).................... ( -6.60, z-score = -1.32, R) >droSim1.chr2R 5508022 54 + 19596830 UUAGUUAAACUUUGCUAAGUAAAGUAUCUUAACUAUUUCAUAGUUUCCGUCUCU .((((((((((((((...)))))))...)))))))................... ( -8.70, z-score = -2.83, R) >droSec1.super_1 4534308 54 + 14215200 UUAGUUAAACUUUGCUAAGUAAAGUAUCUUAACUAUUUCAUAGUUUCCGUCUCU .((((((((((((((...)))))))...)))))))................... ( -8.70, z-score = -2.83, R) >droYak2.chr2R 18962808 54 - 21139217 UUAGUUAAGCUUUACUACACAAAGUAUGUAAACUAUUUCAUAGAUUCUGCCGCU .......(((((((((((.....))).)))))((((...))))........))) ( -5.20, z-score = 0.36, R) >consensus UUAGUUAAACUUUGCUAAGUAAAGUAUCUUAACUAUUUCAUAGUUUCCGUCUCU .((((((((((((((...)))))))...)))))))................... ( -5.60 = -6.48 + 0.88)


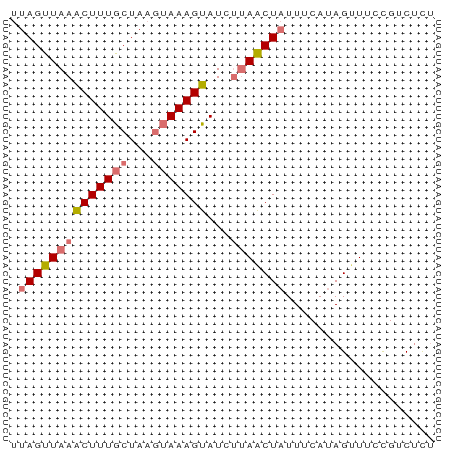
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:14:23 2011