
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,908,908 – 6,908,968 |
| Length | 60 |
| Max. P | 0.694839 |

| Location | 6,908,908 – 6,908,968 |
|---|---|
| Length | 60 |
| Sequences | 4 |
| Columns | 60 |
| Reading direction | reverse |
| Mean pairwise identity | 56.11 |
| Shannon entropy | 0.75376 |
| G+C content | 0.37917 |
| Mean single sequence MFE | -9.83 |
| Consensus MFE | -3.92 |
| Energy contribution | -4.05 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.12 |
| Mean z-score | -0.89 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.694839 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
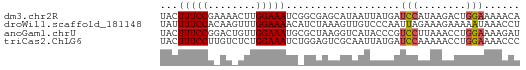
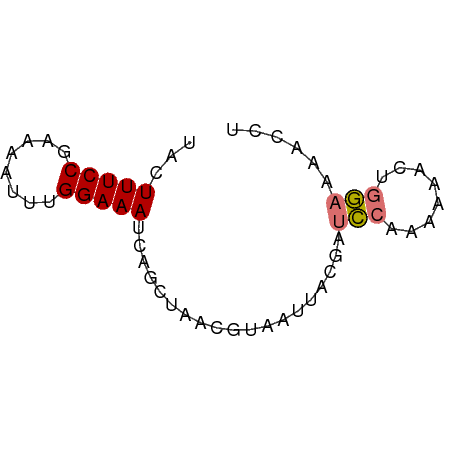
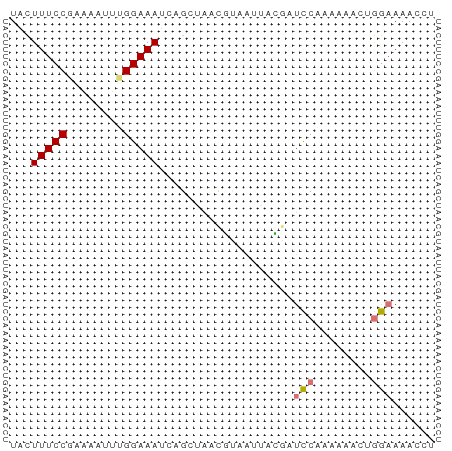
>dm3.chr2R 6908908 60 - 21146708 UACUUUCCGAAAACUUGGAAAUCGGCGAGCAUAAUUAUGAUCCAUAAGACUGGAAAAACA ...(((((((....)))))))...................((((......))))...... ( -8.60, z-score = -0.79, R) >droWil1.scaffold_181148 3777795 60 - 5435427 UAUUUUCCACAAGUUUGGAAAACAUCUAAAGUUGUCCCAAUUAGAAAGAAAAAUAAACCU ........((((.((((((.....)))))).))))......................... ( -7.50, z-score = -1.15, R) >anoGam1.chrU 6326255 60 - 59568033 UACUUUCCGGACUGUUGGAAAUGCGCUAAGGUCAUACCCGUCCUUAAACCUGGAAAAGAU ...(((((((...((.......))..(((((.(......).)))))...))))))).... ( -12.10, z-score = -0.45, R) >triCas2.ChLG6 13069976 60 - 13544221 UACUUUCCUUGUCUCUGGAAAUCUGGAGUCGCAAUUAUGAUCCAAAAACCUGGAAAACCC ..........(.(((..(....)..))).)..........((((......))))...... ( -11.10, z-score = -1.18, R) >consensus UACUUUCCGAAAAUUUGGAAAUCAGCUAACGUAAUUACGAUCCAAAAAACUGGAAAACCU ...(((((........)))))...................(((........)))...... ( -3.92 = -4.05 + 0.13)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:14:06 2011