
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,890,855 – 6,890,923 |
| Length | 68 |
| Max. P | 0.872074 |

| Location | 6,890,855 – 6,890,923 |
|---|---|
| Length | 68 |
| Sequences | 8 |
| Columns | 75 |
| Reading direction | forward |
| Mean pairwise identity | 54.13 |
| Shannon entropy | 0.90620 |
| G+C content | 0.56428 |
| Mean single sequence MFE | -28.36 |
| Consensus MFE | -4.81 |
| Energy contribution | -5.44 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 2.00 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.17 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.872074 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
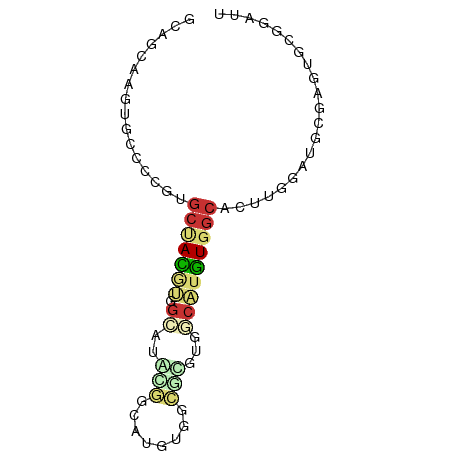
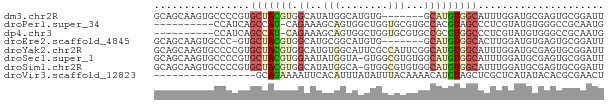
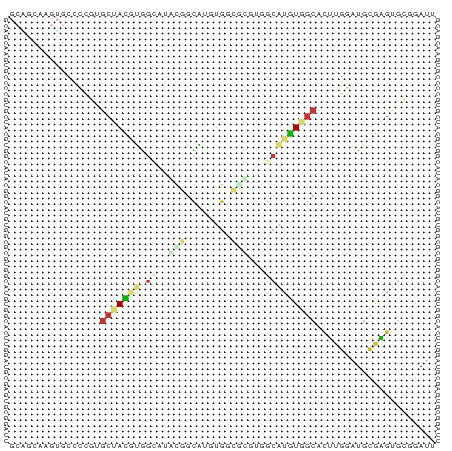
>dm3.chr2R 6890855 68 + 21146708 GCAGCAAGUGCCCCGUGCUACGUGGCAUAUGGCAUGUG-------GCAUGUGGCAUUUGGAUGCGAGUGCGGAUU (((.((((((((.(((((((((((.(....).))))))-------))))).))))))))..)))........... ( -35.70, z-score = -3.69, R) >droPer1.super_34 118635 64 - 916997 ----------CCAUCAGCCAU-CAGAAAGCAGUGGCUGGUGCGUGCCACGUAGCCCUCGUAUGUGGGCCGCAAUG ----------..(((((((((-.........)))))))))(((..(((((((.......)))))))..))).... ( -24.60, z-score = -1.62, R) >dp4.chr3 934311 64 - 19779522 ----------CCAUCAGCCAU-CAGAAAGCAGUGGCUGGUGCGUGCCGCGUGGCCCUCGUAUGUGGGCCGCAAUG ----------.((((((((((-.........))))))))))........(((((((........))))))).... ( -29.50, z-score = -2.36, R) >droEre2.scaffold_4845 3699634 67 - 22589142 GCAGCAAGUGCCC-GUGCUACGUGGCAUGCGGCAUGUG-------GCAUGUGGCACUUGGAUGUGAGUGCGGAUU (((.(((((((((-((((((((((.(....).))))))-------))))).))))))))..)))........... ( -35.30, z-score = -3.37, R) >droYak2.chr2R 18899359 75 - 21139217 GCAGCAAGUGCCCCGUGCUACGUGGCAUGUGGCAUUCGCCAUUCGGCAUGUGGCAUUUGGAUGCGAGUGCGGAUU ...(((..(((.((((((((((((.(..(((((....)))))..).))))))))))..))..)))..)))..... ( -33.50, z-score = -1.68, R) >droSec1.super_1 4472617 74 + 14215200 GCAGCAAGUGCCCCGUGCUACGUGGAAUAUGGUA-GUGGCGUGUGGCAUGUGGCAUUUGGAUGCGAGUGCGGAUU (((.((((((((.(((((((((((..((......-))..))))))))))).))))))))..)))........... ( -29.80, z-score = -1.85, R) >droSim1.chr2R 5445040 74 + 19596830 GCAGCAAGUGCCCCGUGCUACGUGGCAUAUGGCA-GUGGCGUGUGGCAUGUGGCAUUUGGAUGCGAGUGCGGAUU (((.((((((((.(((((((((((.(((......-))).))))))))))).))))))))..)))........... ( -34.70, z-score = -2.15, R) >droVir3.scaffold_12823 286025 58 + 2474545 -----------------GCAGAAAAUUCACAUUUAUAUUUACAAAACAUCUAGCUCGCUCAUAUACACGCGAACU -----------------..................................((.((((..........)))).)) ( -3.80, z-score = -0.66, R) >consensus GCAGCAAGUGCCCCGUGCUACGUGGCAUACGGCAUGUGGCGCGUGGCAUGUGGCACUUGGAUGCGAGUGCGGAUU ................(((((((.....(((.(....).))).....)))))))..................... ( -4.81 = -5.44 + 0.63)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:14:04 2011