
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,886,395 – 6,886,499 |
| Length | 104 |
| Max. P | 0.815553 |

| Location | 6,886,395 – 6,886,499 |
|---|---|
| Length | 104 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 74.10 |
| Shannon entropy | 0.53854 |
| G+C content | 0.41551 |
| Mean single sequence MFE | -25.64 |
| Consensus MFE | -14.23 |
| Energy contribution | -15.17 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.815553 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
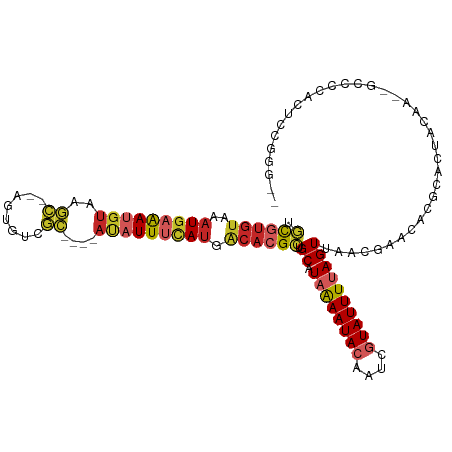
>dm3.chr2R 6886395 104 - 21146708 UGCGUGUAAAUGAAAUGUAAGC--AGUGUCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACACAGGCUACAA--GCUCCACUCCGGG-- .((((((..(((((((((..((--......))----))))))))).)))))).((.((((((((....))))))))(((....)))...........--)).((.....)).-- ( -25.00, z-score = -1.29, R) >droSec1.super_1 4468225 104 - 14215200 UGCGUGUAAAUGAAAUGUAAGC--AGUGUCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACACAGGCUACAA--GCUCCACUCCGGG-- .((((((..(((((((((..((--......))----))))))))).)))))).((.((((((((....))))))))(((....)))...........--)).((.....)).-- ( -25.00, z-score = -1.29, R) >droYak2.chr2R 18894858 104 + 21139217 UGCGUGUAAAUGAAAUGUAAGC--AGUGUCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACACAGGCUACAA--GCUCCACUCCGGC-- .((((((..(((((((((..((--......))----))))))))).)))))).((.((((((((....)))))))).....((......(((.....--)))......))))-- ( -25.40, z-score = -1.30, R) >droEre2.scaffold_4845 3695548 106 + 22589142 UGCGUGUAAAUGAAAUGUAAGC--AGUGUCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACACGGGCUACAAGCGCUCCACUCCGGG-- .((((((..(((((((((..((--......))----))))))))).))))))....((((((((....)))))))).....((.....((((.......)))).....))..-- ( -26.70, z-score = -0.99, R) >droAna3.scaffold_13266 606225 98 - 19884421 UGCGUGUAAAUGAAAUGUAAGC--AGUGUCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACAGGCUACAGGA--UCCCGGG-------- .((((((..(((((((((..((--......))----))))))))).)))))).((.((((((((....))))))))(((....)))..)).......--.......-------- ( -24.20, z-score = -1.29, R) >dp4.chr3 929146 106 + 19779522 UGCGUGUAAAUGAAAUGUAAGC--AGUGCCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUGGCGACCGGGCAGGAGGC--UCCCAGCUGCAGGAC .((((((..(((((((((..((--......))----))))))))).))))))((((((((((((....))))))))(((((.((((........)).--))))))))))).... ( -36.20, z-score = -2.33, R) >droPer1.super_34 113531 106 + 916997 UGCGUGUAAAUGAAAUGUAAGC--AGUGCCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUGGCGACCGGGCAGGAGGC--UCCCAGCUGCAGGAC .((((((..(((((((((..((--......))----))))))))).))))))((((((((((((....))))))))(((((.((((........)).--))))))))))).... ( -36.20, z-score = -2.33, R) >droWil1.scaffold_180697 3211688 97 - 4168966 UGCGUGUAAAUGAAAUGUAAGCAGAGUGUCGC----AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACAAGCAUUCUAA--CCCC----------- .((((((..(((((((((..((........))----))))))))).))))))(((.((((((((....))))))))(((....)))..)))......--....----------- ( -25.30, z-score = -3.16, R) >droMoj3.scaffold_6496 26733066 106 + 26866924 UGUGCGUGCAUGAAACGGCAAC--AGCAGCACAAAAACAUUAUAUAGCAGGAUGCGUUAAAUAAAAUCAUAUUUUAGUGAACGAACAAGAACUCAGAUAAGCCGUCUG------ (((((.(((.((.........)--))))))))).....................((((...((((((...))))))...))))..........(((((.....)))))------ ( -16.30, z-score = 1.14, R) >droGri2.scaffold_15245 6354994 105 - 18325388 UAUGUCCAAAUAUGACAUACAU--AUAAGCAU----ACAUUUCAAGCCAAGUUGCGUUGAAUACAAUCAUAUUUUAGUUAACAAGCAGCCACAAGAA-GGAUAAGCUGCUGC-- ((((((.......))))))...--....((..----....(((((((......)).)))))......................((((((........-......))))))))-- ( -16.14, z-score = 0.41, R) >consensus UGCGUGUAAAUGAAAUGUAAGC__AGUGUCGC____AUAUUUCAUGACACGCUGCAUAAAAUACAAUCGUAUUUUAGUUAACGAACACGCACUACAA__GCCCCACUCCGGG__ .((((((..(((((((((..((........))....))))))))).)))))).((.((((((((....)))))))))).................................... (-14.23 = -15.17 + 0.94)
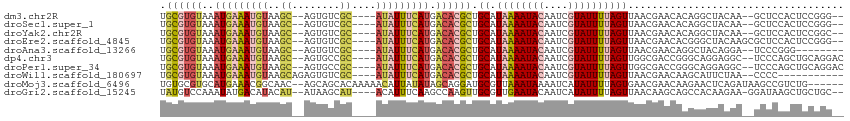


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:14:03 2011