
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,862,198 – 6,862,313 |
| Length | 115 |
| Max. P | 0.530588 |

| Location | 6,862,198 – 6,862,313 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.43 |
| Shannon entropy | 0.27718 |
| G+C content | 0.46133 |
| Mean single sequence MFE | -35.75 |
| Consensus MFE | -19.41 |
| Energy contribution | -20.75 |
| Covariance contribution | 1.34 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.530588 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6862198 115 - 21146708 AACUUU----UGGCGUCUUUAAGCCAAACUCGCUUUUUAUAAAUCCUCUUUGGGAAUUCCGAUGG-GGGAUUAACGGCUCUCGCAGAGGAGACAUCUGCCGAUUGCCACAAAAGUUGCCA ....((----((((........))))))...((((((....((((((((.(.((....)).).))-))))))...(((..((((((((....).)))).)))..)))..))))))..... ( -39.80, z-score = -3.20, R) >droSim1.chr2R 5419214 115 - 19596830 AACUUU----UGGCGUCUUUAAGCCAAACUCGCUUUUUAUAAAUCCUCUUUGGGAAUUCCGAUGG-GGGAUUAACGGCUCUCGCAGCGGAGACAUCUGCCGAUUGCCACAAAAGUUGCCA ....((----((((........))))))...((((((....((((((((.(.((....)).).))-))))))...(((..(((..(((((....))))))))..)))..))))))..... ( -37.50, z-score = -2.45, R) >droSec1.super_1 4444741 115 - 14215200 AACUUU----UGGCGUCUUUAAGCCCAACUCGCUUUUUAUAAAUCCUCUUUGGGAAAUCCGAUGU-GGGAUUAACGGCUUUCGCAGCGGAGACAUCUGCCGAUUGCCACAAAAGUUGCCA ......----(((((.((((...((((..(((..((((.((((.....)))).))))..)))..)-)))......(((..(((..(((((....))))))))..)))...)))).))))) ( -34.70, z-score = -1.99, R) >droYak2.chr2R 18870347 115 + 21139217 AACUUU----UGGCGUCUUUAUGCCAAACUCGCUUUUUAUAAGUCCUCUUCAGGCAUUCCGAUGG-GGGAUUUGCGGCUCUCGCAGCGGAGACAUCUGCCGAUUGCCACAAAAGUUGCCA ....((----((((((....))))))))...((((((..((((((((((...((....))...))-)))))))).(((..(((..(((((....))))))))..)))..))))))..... ( -38.00, z-score = -1.97, R) >droEre2.scaffold_4845 3669049 115 + 22589142 AACUUU----GGGCGUCUUUAAGCCAAACUCGCUUUUUAUAAAUCCUCUUUUGGAAUUCCAAUGG-GGGAUUAACGGCUCUCGCAGCGGAGACAUCUGCCGAUUGCCACAAAAGUUGCCA ......----.((((.((((((((.......))))......((((((((.((((....)))).))-))))))...(((..(((..(((((....))))))))..)))...)))).)))). ( -38.90, z-score = -3.13, R) >droAna3.scaffold_13266 582414 119 - 19884421 GACUUUAUCCUAAAGUGCCUAAUCCUAACUUGCUCCGGUUAUAUCCUCUUUA-AAAUUCCGGUGGCGAGAUUAACAGUUCCAUGGGCGAAGACAUCUGUCAAUUGCCACAAAAGUUGCAA ((((((......................((((((((((..............-.....)))).)))))).............(((((((.(((....)))..))))).)))))))).... ( -25.61, z-score = -0.73, R) >consensus AACUUU____UGGCGUCUUUAAGCCAAACUCGCUUUUUAUAAAUCCUCUUUGGGAAUUCCGAUGG_GGGAUUAACGGCUCUCGCAGCGGAGACAUCUGCCGAUUGCCACAAAAGUUGCCA ((((((.....(((........)))................((((((((...((....))...)).))))))...(((..(((..(((((....))))))))..)))...)))))).... (-19.41 = -20.75 + 1.34)

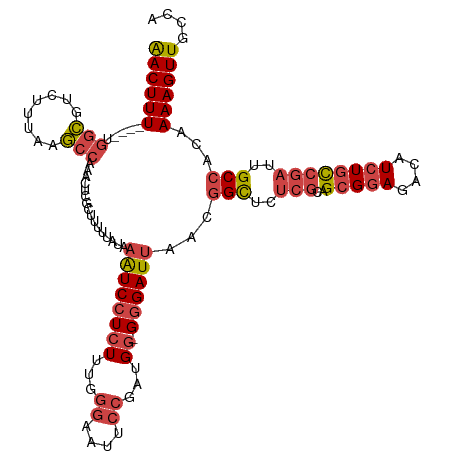
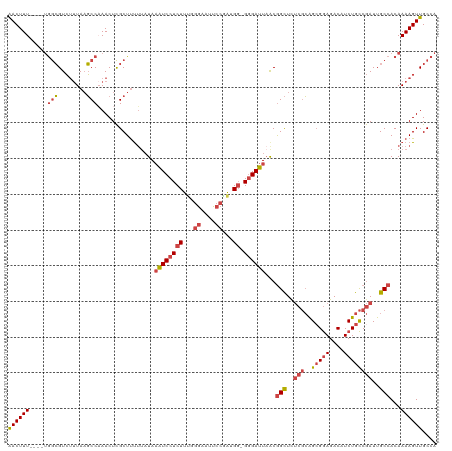
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:13:59 2011