
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,830,012 – 6,830,110 |
| Length | 98 |
| Max. P | 0.992269 |

| Location | 6,830,012 – 6,830,110 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 82.38 |
| Shannon entropy | 0.32767 |
| G+C content | 0.49662 |
| Mean single sequence MFE | -24.52 |
| Consensus MFE | -16.76 |
| Energy contribution | -16.96 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.27 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.53 |
| SVM RNA-class probability | 0.992269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6830012 98 + 21146708 GAAUGGCGCCCACCCUUUU------GUACCCAUACUGCCCAUCUCUCUCGGCAUUGUUCUACC--CAUCUCUUUCUCAUAAAGAGACUGCAGUUAUGCAUUGCUGU ((((((.((..........------...........)))))).))...(((((.(((...((.--(((((((((.....))))))).))..))...))).))))). ( -19.30, z-score = -1.30, R) >droSim1.chr2R 5398693 98 + 19596830 GAAUGGCGCCCUCCCUUUU------GUGCCCAUUCUACCCGUCUCUCUCGGCAUUGUUCUACC--CGUCUCUUUCUCAUAAAGAGACUGCAGUUAUGCAUUGCUGU ((((((.((.(........------).)))))))).............(((((.(((...((.--.((((((((.....))))))))....))...))).))))). ( -27.50, z-score = -4.26, R) >droSec1.super_1 4420380 97 + 14215200 GAAUGGCGCCCUCCCUUUU------GUGCCCAUUCUACCCGUCUC-CUCGGCAUUGUUCUACC--CGUCCCUUUCUCAUAAAGAGACUGCAGUUAUGCAUUGCUGU ((((((.((.(........------).))))))))..........-..(((((.(((...((.--.(((.((((.....)))).)))....))...))).))))). ( -22.70, z-score = -2.63, R) >droYak2.chr2R 18844570 98 - 21139217 GAAUGGCGCCCUCCCUUUU------GUGCCCAUUCUACCCAUCUCUCUCGGCAUUGUUCUACC--CGUCUCUUUCUCAUAAAGAGACUGCAGUUAUGCAUUGCUGU ((((((.((.(........------).)))))))).............(((((.(((...((.--.((((((((.....))))))))....))...))).))))). ( -27.50, z-score = -4.40, R) >droEre2.scaffold_4845 3644503 97 - 22589142 GAAUGGCGCCCUCC-UUUU------GUGCCCACUCUACCCAUCUCUCUCGGCAUUGUUCUACC--CGUCUCUUUCUCAUAAAGAGACUGCAGUUAUGCAUUGCUGU ....(((((.....-....------)))))..................(((((.(((...((.--.((((((((.....))))))))....))...))).))))). ( -23.50, z-score = -2.82, R) >droAna3.scaffold_13266 560117 102 + 19884421 ----GCCAUCCUCCUCUAGAAGGGAGUGCCUUCUCUUUCCCCUGCACUAGUCCCUUCUCUUUCACUGUUUUUGUCUCACACGGAGACUGCAGUUAUGCAUUGCUGU ----((...........((((((((((((..............))))...)))))))).....(((((....(((((.....))))).)))))...))........ ( -26.64, z-score = -1.85, R) >consensus GAAUGGCGCCCUCCCUUUU______GUGCCCAUUCUACCCAUCUCUCUCGGCAUUGUUCUACC__CGUCUCUUUCUCAUAAAGAGACUGCAGUUAUGCAUUGCUGU ....(((((................)))))..................(((((.............((((((((.....))))))))((((....)))).))))). (-16.76 = -16.96 + 0.20)



| Location | 6,830,012 – 6,830,110 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 82.38 |
| Shannon entropy | 0.32767 |
| G+C content | 0.49662 |
| Mean single sequence MFE | -27.06 |
| Consensus MFE | -15.98 |
| Energy contribution | -16.52 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.642794 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6830012 98 - 21146708 ACAGCAAUGCAUAACUGCAGUCUCUUUAUGAGAAAGAGAUG--GGUAGAACAAUGCCGAGAGAGAUGGGCAGUAUGGGUAC------AAAAGGGUGGGCGCCAUUC ...((.((((....((((.((((((((.....)))))))).--.)))).....((((.(......).))))))))..))..------....((((((...)))))) ( -26.30, z-score = -1.51, R) >droSim1.chr2R 5398693 98 - 19596830 ACAGCAAUGCAUAACUGCAGUCUCUUUAUGAGAAAGAGACG--GGUAGAACAAUGCCGAGAGAGACGGGUAGAAUGGGCAC------AAAAGGGAGGGCGCCAUUC .......(((....((((.((((((((.....)))))))).--.)))).......(((.......))))))((((((((.(------........).)).)))))) ( -26.80, z-score = -2.23, R) >droSec1.super_1 4420380 97 - 14215200 ACAGCAAUGCAUAACUGCAGUCUCUUUAUGAGAAAGGGACG--GGUAGAACAAUGCCGAG-GAGACGGGUAGAAUGGGCAC------AAAAGGGAGGGCGCCAUUC ..............((((.((((((((.....)))))))).--.)))).....((((..(-....).))))((((((((.(------........).)).)))))) ( -28.70, z-score = -2.76, R) >droYak2.chr2R 18844570 98 + 21139217 ACAGCAAUGCAUAACUGCAGUCUCUUUAUGAGAAAGAGACG--GGUAGAACAAUGCCGAGAGAGAUGGGUAGAAUGGGCAC------AAAAGGGAGGGCGCCAUUC ..............((((.((((((((.....)))))))).--.)))).....((((.(......).))))((((((((.(------........).)).)))))) ( -27.30, z-score = -2.43, R) >droEre2.scaffold_4845 3644503 97 + 22589142 ACAGCAAUGCAUAACUGCAGUCUCUUUAUGAGAAAGAGACG--GGUAGAACAAUGCCGAGAGAGAUGGGUAGAGUGGGCAC------AAAA-GGAGGGCGCCAUUC ..............((((.((((((((.....)))))))).--.)))).....((((.(......).))))((((((((.(------....-....))).)))))) ( -27.10, z-score = -2.23, R) >droAna3.scaffold_13266 560117 102 - 19884421 ACAGCAAUGCAUAACUGCAGUCUCCGUGUGAGACAAAAACAGUGAAAGAGAAGGGACUAGUGCAGGGGAAAGAGAAGGCACUCCCUUCUAGAGGAGGAUGGC---- ...((..((((....)))).(((((.(((.........))).......((((((((...((((..............))))))))))))...)))))...))---- ( -26.14, z-score = -0.41, R) >consensus ACAGCAAUGCAUAACUGCAGUCUCUUUAUGAGAAAGAGACG__GGUAGAACAAUGCCGAGAGAGAUGGGUAGAAUGGGCAC______AAAAGGGAGGGCGCCAUUC .......(((....((((.((((((((.....))))))))...((((......))))...........)))).....))).......................... (-15.98 = -16.52 + 0.53)


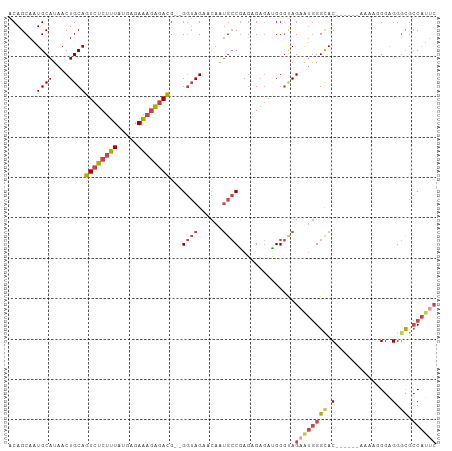
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:13:47 2011