
| Sequence ID | dm3.chr2R |
|---|---|
| Location | 6,806,829 – 6,806,925 |
| Length | 96 |
| Max. P | 0.855708 |

| Location | 6,806,829 – 6,806,925 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 73.77 |
| Shannon entropy | 0.50349 |
| G+C content | 0.43288 |
| Mean single sequence MFE | -30.61 |
| Consensus MFE | -11.21 |
| Energy contribution | -12.15 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.855708 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2R 6806829 96 + 21146708 GCUGUCAAUAAUUUAUAGGCCCUGAGGCAAAAACUUUACUGCCAAA-GUGCCGUAAAUUGGCAG---CUC-GCA-GUUGCUUUUGUGGUU-CUAUGUCAGUGU------------- (((((((((...((((.(((.((..((((..........))))..)-).)))))))))))))))---)((-(((-(......))))))..-............------------- ( -28.00, z-score = -1.42, R) >droAna3.scaffold_13266 534144 94 + 19884421 GCUGUCAAUAAUUUAUAGGCAA-GAGGCAAAAACUUUACGGCCAAA-GUGCCGUAAAUUGGCAG---CUCCGAA-GUUGUUUCUGUGGCG-GUGUCUGCAU--------------- (((((((.((((((((.((((.-..(((............)))...-.))))))))))))((((---((....)-))))).....)))))-))........--------------- ( -26.20, z-score = -0.62, R) >droEre2.scaffold_4845 3621573 96 - 22589142 GCUGUCAAUAAUUUAUAGGCGCUGAGGCAAAAACUUUACUGCCAAA-GUGCCGUAAAUUGGCAG---CUC-GCA-GUUGCCUUUGUGGUU-CUAUGUGUGACU------------- (((((((((...((((.((((((..((((..........))))..)-)))))))))))))))))---).(-(((-...(((.....))).-...)))).....------------- ( -35.00, z-score = -3.31, R) >droYak2.chr2R 18818263 96 - 21139217 GCUGUCAAUAAUUUAUAGGCGCUGAGGCAAAAACUUUACUGCCAAA-GUGCCGUAAAUUGGCAG---CUC-GCA-GUUGCCUUUGUGGUU-CUAUGUGUAUGU------------- (((((((((...((((.((((((..((((..........))))..)-)))))))))))))))))---).(-(((-...(((.....))).-...)))).....------------- ( -35.00, z-score = -3.46, R) >droSec1.super_1 4396134 92 + 14215200 GCUGUCAAUAAUUUAUAGGCGCUGAGGCAAAAACUUUACUGCCAAA-GUGCCGUAAAUUGGCAG---CUC-GCA-GUUGCUUUUGUGGUU-CUAUGUAA----------------- (((((((((...((((.((((((..((((..........))))..)-)))))))))))))))))---)((-(((-(......))))))..-........----------------- ( -32.40, z-score = -3.12, R) >droSim1.chr2R 5382755 94 + 19596830 GCUGUCAAUAAUUUAUAGGCGCUGAGGCAAAAACUUUACUGCCAAA-GUGCCGUAAAUUGGCAG---CUC-GCA-GUUGCUUUUGUGGUU-CUAUGUAACU--------------- (((((((((...((((.((((((..((((..........))))..)-)))))))))))))))))---)((-(((-(......))))))..-..........--------------- ( -32.40, z-score = -2.98, R) >droWil1.scaffold_180697 3120960 97 + 4168966 GCUGUCAAUAAUUUAUAUGUG-------AAAGUUGCUACGACCAAAAGUGCCGUAAAUUGGCAA--------AAAGUUGCCAGUGUACCAAGAACAUUCAUACCCCCGAAGA---- .................((((-------((.(((((.((........)))).(((.((((((((--------....)))))))).)))....))).))))))..........---- ( -18.80, z-score = -1.53, R) >droMoj3.scaffold_6496 26673269 109 - 26866924 GCUGUCAAUAAUUUAUUCGCC----GGCAAAA-CUCUACAGCAAAA-GUGCCGUAAAUUGGCAGUGGCUGUUUACGUUGUUGUUGUUGUA-GUUGUUGCAGCUGCUACUACAAGCA ((((.((((((((....((.(----(((((..-....(((((...(-.(((((.....))))).).))))).....)))))).))....)-))))))))))).(((......))). ( -37.10, z-score = -2.25, R) >consensus GCUGUCAAUAAUUUAUAGGCGCUGAGGCAAAAACUUUACUGCCAAA_GUGCCGUAAAUUGGCAG___CUC_GCA_GUUGCUUUUGUGGUU_CUAUGUGAAU_U_____________ .((((((((...((((.((((....(((............))).....)))))))))))))))).................................................... (-11.21 = -12.15 + 0.94)

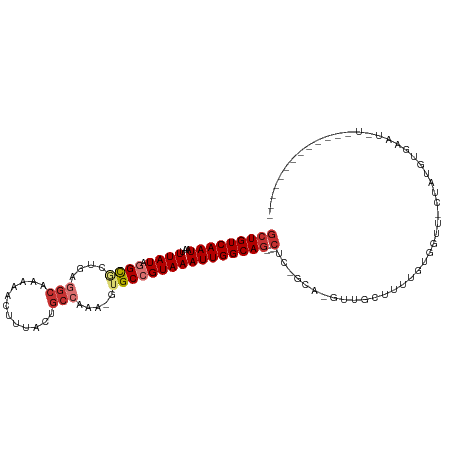

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 22:13:44 2011